Bacterial Mercury Methylation Enhanced by Mixed Thiols
02/01/2023
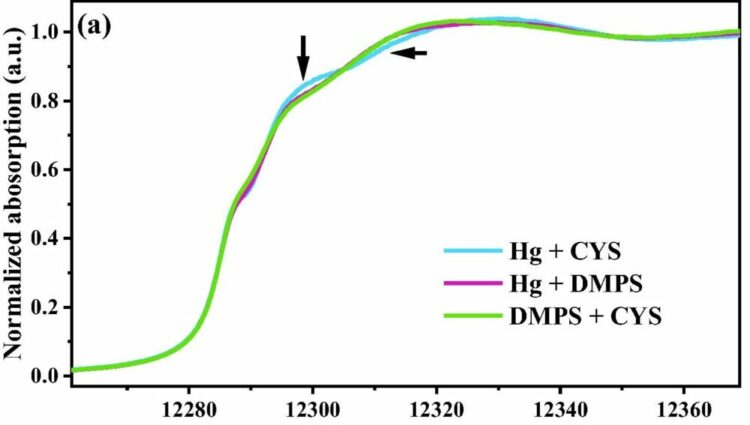
Mercury transformation in the environment. X-ray absorption spectroscopy helped determine whether the coordination environments of mercury (Hg) complexed with different thiol compounds (cysteine, DMPS, DMSA, or a combination) affect Hg methylation. Differences among the spectra indicate a mixture of Hg(II)-thiolate species with coordination numbers greater than 2, which may facilitate Hg methylation. [Reprinted from Geochimica et Cosmochimica Acta, Vol 342, X. Liang et al., High methylation potential of mercury complexed with mixed thiolate ligands by Geobacter sulfurreducens PCA, Pages No. 74-83, Copyright 2023, with permission from Elsevier.]
The Science
Inorganic mercury (Hg) released into the environment by humans is converted to the toxin monomethylmercury (MgHg) by certain anaerobic microorganisms. MeHg is present in waterways and bioaccumulates through aquatic food chains, posing a risk to wildlife and human health. Thus, it is important to understand biological control of environmental Hg transformation. Researchers at Oak Ridge National Laboratory (ORNL) and the Stanford Synchrotron Radiation Lightsource (SSRL) investigated how the presence of different low-molecular-weight thiol compounds, known to either inhibit or enhance Hg methylation, influence rates and extent of Hg methylation by the model methylating bacterium Geobacter sulfurreducens PCA (PCA).
The Impact
Results indicate that the effects of thiols on Hg methylation are more complex than previously known. Mercury complexed individually with the thiols cysteine, dithiol 2,3-dimercaptopropanesulfonate (DMPS), or 2,3-dimercaptosuccinic acid (DMSA) at low concentrations found in the environment strongly inhibited Hg methylation. Yet a mixture of cysteine with either DMSA or DMPS increased methylation rates by 1.5x-3.5x over the no-thiol control.
Spectroscopic analyses indicated the formation of mixed Hg-thiolate complexes with coordination numbers of 3 or 4, which likely facilitated exchange of Hg with cells and its uptake and internal transfer to the proteins that carry out methylation. This foundational study underscores the need for detailed spectroscopic studies of environmental pollutants and biological controls that could act as natural pollutant remediation processes.
The Summary
Extended X-ray absorption fine structure (EXAFS) and X-ray absorption near edge structure (XANES) spectroscopy were performed at SSRL beamline 7-3 on mixtures of low-molecular-weight thiols complexed with Hg to determine potential mechanisms of complexation. Spectroscopy data were complemented by density functional theory calculations also conducted at SSRL, as well as ultraviolet-visible (UV-vis) and Raman spectroscopies at ORNL.
Related Links
References
Liang et al. 2023. High Methylation Potential of Mercury Complexed with Mixed Thiolate Ligands by Geobacter sulfurreducens PCA. Geochimica et Cosmochimica Acta 342, 74–83. [DOE:10.1016/j.gca.2022.12.008]
