Highlights
Research utilizing BER-supported structural biology and imaging resources at DOE user facilities
| Date | Title | Resource | Technique | Facility | Research Theme | ||||
|---|---|---|---|---|---|---|---|---|---|
| 03/07/2024 | Multimodal X-Ray Imaging Technique for Single Cells | Laboratory for BioMolecular Structure | Hard X-ray Tomography, X-ray Fluorescence Imaging | National Synchrotron Light Source II | Cell and Tissue Structure | Understanding the inner workings of cells and their interactions at such detailed levels can transform the approach to various fields, particularly in identifying how pathogens interact with host cells. This multimodal imaging approach provides insights that could improve agricultural practices by studying plant–pathogen interactions and enhance drug delivery systems. These insights could lead to better strategies for improving crop resilience and combating infectious diseases. | Scientists from DOE’s Brookhaven National Laboratory used X-ray computed tomography (XCT) and X-ray fluorescence (XRF) microscopy to develop a novel multimodal method to noninvasively image a single cell, achieving nanometer resolution without damaging delicate organelles. The method demonstrates the feasibility and broad applicability of using X-rays to understand cellular structures and the roles of chemical elements and related proteins in signaling and other biological processes. | Cells from the human embryonic kidney (HEK) 293 line were chemically preserved using paraformaldehyde to maintain cellular structure. They were then rapidly frozen, transferred to liquid nitrogen, and freeze-dried before being placed under a microscope to locate and label them for targeted imaging. Standardized sample holders were created for use on multiple pieces of equipment to ensure cells could be reliably held in place for multiple measurements. The team used two imaging techniques found at NSLS-II: XCT and XRF microscopy. The Full Field X-ray Imaging (FXI) beamline was used to collect XCT data. The technique uses X-rays to show a cross-section of a solid sample, providing information about a cell’s physical structure. This is similar to computed tomography scans in medical imaging. The researchers also used the Submicron Resolution X-ray Spectroscopy (SRX) beamline to collect XRF microscopy data, which provides information about the distribution of chemical elements within a cell. XRF directs high-energy X-rays at a sample, exciting the material and causing it to emit X-ray fluorescence. This X-ray emission has a unique signature, enabling scientists to determine exactly which elements comprise the sample and how they are distributed to fulfill their biological functions. Achieving high-resolution fluorescence maps of multiple cells can be very time consuming, even for two-dimensional images. The XCT technique, which produces three-dimensional images, can help speed the process. The information can help guide two-dimensional fluorescence measurements to specific locations of interest. This saves time, increases throughput, and reduces the length of time samples are exposed to X-rays, which mitigates potential damage to fragile cells. | |
| 08/01/2024 | Molecular Alcove Required for Anaerobic Carbon Fixation | Structural Molecular Biology Resource | Synchrotron Infrared Hyperspectral Imaging, X-ray Absorption and Emission Spectroscopy | Stanford Synchrotron Radiation Lightsource | Molecular Structure | The alcove formed by the described residues augments and tunes the gas-bonding properties of the active-site Ni2-iron-sulfur metallo-cofactor. Mutation of these residues presents new ways in which the carbon-fixing capabilities of the Wood-Ljungdahl pathway can be enhanced. These residues are now considered integral to ACS functioning to the extent that they should be considered part of the active site, despite not being directly bound to the Ni2-iron-sulfur metallo-cofactor. Similar, yet-undescribed, alcoves likely exist in other gas-handling metalloenzymes. These should be sought out to improve understanding of small-gas metabolism, which underpins all life. | The anaerobic Wood-Ljungdahl pathway is one of seven known carbon fixation pathways, and the only one which conserves energy through net generation of adenosine triphosphate (ATP). At the heart of the pathway are two proteins—carbon monoxide dehydrogenase (CODH) and acetyl-CoA synthase (ACS)—which turn carbon dioxide (CO2) into acetate through a series of organometallic intermediates. In the pathway, ACS accepts CO, produced from CO2 by CODH, and a methyl group from a cobalamin-dependent methyl transferase to form acetyl-CoA at its active site, which is composed of an iron-sulfur cluster adjacent to two nickel (Ni) atoms. CO, a critical intermediate of the pathway, travels to the active site of ACS from CODH through a series of internal gas channels, allowing the organism to tightly control this otherwise toxic substance. In this study, researchers use X-ray absorption spectroscopy and other techniques to demonstrate that five highly conserved hydrophobic residues create a pocket, or alcove, just above the ACS active site where CO enters and is concentrated, favoring its binding to the active site’s catalytic Ni. There, it is combined with the methyl group and CoA to yield the acetyl-CoA product. Through the use of in vitro and in vivo growth, kinetic, spectroscopic, and binding experiments, one of the five alcove residues, F229, was found to play a pivotal role in coordinating CO binding to the ACS active site in a size-dependent manner. Mutation of this residue to the much smaller residue alanine was found to decrease the CO affinity of the active site 30-fold, abolishing the ability of the host organism to grow autotrophically on CO2. In contrast, mutation of F229 to the much larger residue tryptophan enhanced CO affinity 80-fold. | ||
| 10/01/2024 | X-Ray Spectroscopy Reveals Potassium Transport by Ectomycorrhizal Fungi | Structural Molecular Biology Resource | X-ray Absorption and Emission Spectroscopy, X-ray Fluorescence Imaging | Stanford Synchrotron Radiation Lightsource | Chemical and Elemental Information | This research reveals that XRF imaging and XANES spectroscopy can be used to identify the various forms of K accumulating in biological systems, something which is understudied, and shows promise in deepening understanding of the role ECM fungi play in K acquisition and transport. More established and dense ECM have several, more complicated K chemistries compared to younger individual hyphae. This suggests that older versus newer hyphae have different roles in the sourcing, transport, storage, and transfer of K between ECM and host plant symbiotic partners. The ECM fungus Paxillus ammoniavirescens is considered a generalist that associates with many tree species, and has recently been observed to improve K nutrition and improve salinity stress in pine trees. Improving understanding of K uptake, storage, and transport mechanisms by this fungi is therefore applicable to a broad range of ECM and host plants. | Ectomycorrhizal (ECM) fungi are symbiotic microbes essential to many plants, facilitating nutrient transfer via root colonization. Critical plant nutrients, such as carbon (C), nitrogen (N), phosphorus (P), and potassium (K), are transferred through this mutualistic partnership. These nutrients also play a role in tolerance to environmental stressors and rhizosphere nutrient cycling. Compared to N and P, the uptake, transport to plants, and storage of K by ECM is poorly studied. However, K is pivotal to several plant processes, such as enzyme activation (impacting ATP production), osmoregulation, and disease resistance. Despite this, K is limited in most environments due to its strong association with minerals. Therefore, improving and maintaining sufficient K concentrations in plants is essential for plant efficiency and vitality in a diverse range of ecosystems. Researchers used a combination of potassium K-edge X-ray fluorescence (XRF) imaging and X-ray absorption near-edge structure (XANES) spectroscopy on P. ammoniavirescens to investigate the distribution of K chemistries across hyphal biomass. The approach revealed the presence of several K counter-ions to carboxylic acids. This may suggest that, besides their direct transfer to colonized roots, K ions can also be involved in the production of compounds necessary for sourcing nutrients from their surrounding environment by ECM fungi. Additionally, this work reveals that XANES spectroscopy can be used to identify the various forms of K accumulating in biological systems. | P. ammoniavirescens was grown in the absence of a plant host on a solid modified Melin-Norkans medium at 26°C for 3 weeks. Dense hyphal biomass formed on the surface of the media. To prepare hyphal samples for synchrotron analyses, X-ray-compatible tape was placed on the dense biomass, maintaining a known spatial orientation of the mycelium. Researchers mapped the distribution of K, S, and P along hyphae and observed the accumulation of different K chemistries in different parts of the mycelium, as well as colocations of K and P. K-nitrate (KNO3), K-C-O [such as K-tartrate K2(C4H4O6) and K-oxalate (K2C2O4)], K-S, and K-P compounds were all present in P. ammoniavirescens. In particular, K-C-O compounds were present as hotspots either on the outside of younger mycelia or within traceable hypha in the older mycelium. This is likely associated with the production and exudation of low-molecular-weight organic acids within younger mycelia, but storage and transport in older mycelia. | |
| 01/29/2024 | X-ray Spectroscopy Reveals Potassium-Bonding Environment | Structural Molecular Biology Resource | X-ray Absorption and Emission Spectroscopy | Environmental Molecular Sciences Laboratory, Stanford Synchrotron Radiation Lightsource | Molecular Structure | This research shows that the bonding environment of potassium in different salts of organic acids has different detectable signatures using XAS. The resulting spectra can inform whether potassium is associated with carbon, nitrogen, or oxygen. This will enable future research to fingerprint the type of organic compound bonded to K in complex biological and environmental samples; something that was not previously known to be possible. A further impact of this research is that characterization of these signatures will enable future studies to spatially distinguish among potassium organic molecules in natural soils. | Potassium (K) is an essential element for plant growth. Soils can contain up to 3% K by weight, but the majority is structurally bound within minerals and considered non-bioavailable. Therefore, most environments are K-limited, even when supplied with fertilizer. Weathering of K-rich minerals has the capacity to increase bioavailable K in the rhizosphere, but abiotic mineral weathering is too slow to support crop growth. Fortunately, several microbial species use mechanical and chemical processes to weather mineral surfaces and release critical nutrients to the local environment. However, the molecular mechanisms underpinning the biochemical processes involved in microbial nutrient sourcing from minerals are poorly understood. To better understand how microbes source K from minerals, scientists at the Stanford Synchrotron Radiation Lightsource performed X-ray absorption spectroscopy (XAS) on K-organic salts, including acetate, citrate, nitrate, oxalate, and tartrate, which are frequently observed as acids secreted by soil microbes. Results showed that XAS spectra are associated with extended organic salt ligands, and unique and distinct spectral features are associated with second-shell nitrogen compared to carbon. The improved understanding of K bonding environments with organic compounds provides an important toolkit to understand how K is transformed by microbial processes and made bioavailable for plant uptake. | K-organic salts displayed feature-rich XAS spectra, each demonstrating numerous unique features spanning 13 eV, despite similar first shell bonding environments. To identify the electronic transitions that give rise to some of the unique spectral features in the organic salts, researchers used computational tools including molecular dynamics (MD), time-dependent density functional theory (TD-DFT), and full multiple scattering (FMS) in OCEAN, to simulate the experimental spectra. | |
| 04/12/2024 | Iron Availability Tied to Carbon Metabolism in Cyanobacteria | Center for BioMolecular Structure, Structural Molecular Biology Resource | Solution X-ray Scattering, X-ray Absorption and Emission Spectroscopy, X-ray Macromolecular Crystallography | National Synchrotron Light Source II, Stanford Synchrotron Radiation Lightsource | Molecular Structure | The discovery of Dri1 and its role in the regulation of photosynthesis and respiration in cyanobacteria provides new insight into how CO2 removal via photosynthesis is dependent on iron availability. Since iron is often limited in marine environments where many species of cyanobacteria are located, the discovery of this regulatory mechanism offers an explanation into how these abundant organisms are able to thrive under these nutrient limited conditions, and offers insight into how iron homeostasis and carbon metabolism are interlinked. | Cyanobacteria have photosynthetic and respiratory pathways that are physically interlinked, whereas in plants these pathways are separated into different organelles. In cyanobacteria, multiple enzymatic complexes, including the well-characterized type I and type II NADH dehydrogenases and succinate dehydrogenase (Sdh), contribute electrons into both the photosynthetic and respiratory pathways through their participation in the redox poising of the plastoquinone pool. The enzymatic complexes share iron-dependent electron carriers between the photosynthetic and respiratory electron transfer chains. This adds an extra regulatory burden on the cell, which needs to tightly coordinate iron homeostasis with photosynthesis and respiration. Scientists from Brookhaven National Laboratory, SLAC National Accelerator Laboratory, and Lawrence Berkeley National Laboratory have discovered a new highly conserved protein from the cyanobacterium Synechocystis sp. PCC 6803 that jointly regulates these two pathways with iron homeostasis. The protein, called Dri1 (for Domain related to Iron 1), uses a never-before-seen heme-binding structural motif. Using macromolecular crystallography, small-angle X-ray scattering, and Fe K-edge X-ray absorption spectroscopy, researchers were able to characterize Dri1, finding a novel Zinc-mirror heme-binding site within. Dri1 was found to regulate Sdh, which requires many iron-containing cofactors to function. Under conditions of iron limitation, monomeric Dri1 binds to a subunit of Sdh, preventing the formation of the functional protein complex and inhibiting its contribution to the electron transfer chain. When iron isn’t limiting, Dri1 binds iron in the form of a heme-Dri1 dimer; when Zn is also bound, it tightly coordinates the heme, preventing its release back into the cytosol and inhibiting the interaction between Dri1and Sdh. | This study presents a new protein, Dri1, conserved in cyanobacteria, which links iron homeostasis with carbon metabolism. The structure of this protein reveals a novel Zn-mirror heme-binding motif, wherein Zn binding aligns two histidine residues to tightly coordinate the heme iron. When heme is bound, Dri1 is no longer able to interact with the succinate dehydrogenase (Sdh) subunit SdhB. This allows formation of the fully functional Sdh complex, which is then able to participate as an electron donor for the photosynthetic and respiratory metabolic pathways. | |
| 06/27/2024 | Role of Enigmatic "Red Body" in a Biofuel-Candidate Alga | Berkeley Synchrotron Infrared Structural Biology Imaging Program | Synchrotron Infrared Hyperspectral Imaging | Advanced Light Source | Chemical and Elemental Information | As the effects of climate change continue to grow, scientists face mounting calls to deliver alternative fuels with carbon-neutral emissions when burned. Eustigmatophytes, a group of single-celled algae found in freshwater, marine, and terrestrial environments, could offer an opportunity to produce new biofuels. However, a limited understanding of the life cycle and cell biology of the model species N. oceanica has restricted researchers’ ability to draw conclusions that could otherwise inform cultivation and genetic engineering directions of additional algae species considered to be good candidates for producing biofuels and/or sequestering carbon. This work represents a new biological link between molecular and large-scale processes in cellular systems. | Scientists have long searched for new ways to make fuel and have consequently studied algae that naturally produce and retain large amounts of fatty molecules. Yet questions remain regarding the basic life cycles of algal species of interest, such as Nannochloropsis oceanica and other Eustigmatophytes. The marine nanophytoplankton N. oceanica possess an enigmatic structural feature called the “red body.” Now, Fourier transform infrared spectroscopy has revealed that the globular red body contains an accumulation of antioxidant carotenoids, responsible for the red color, and large quantities of long-chain aliphatic lipids, a type of fatty molecule. In the same study, ultra-performance liquid chromatography coupled with high-resolution mass spectrometry detected a C32 alkyl diol, a potential precursor of the material algaenan, which is a recalcitrant cell wall polymer produced by some green algae. Transmission electron microscope imaging and 3D cryo-tomography indicated the red body is a membrane-bound organelle that likely facilitates the transport of key molecules needed for cell wall construction, supporting the ability of N. oceanica to rapidly divide into two, four, or even eight daughter cells every 24 hours. Without the organelle, N. oceanica could face challenges transporting the hydrophobic molecules, which include the C32 alkyl diol, through its water-based interior. This study involved collaboration among researchers from the University of California-Berkeley, the Lawrence Berkeley National Laboratory (LBNL), and the University of Copenhagen in Denmark. LBNL is home to the Berkeley Synchrotron Infrared Structural Biology (BSISB) program, funded by DOE’s Biological and Environmental Research program, and the Advanced Light Source, a DOE Office of Science user facility. | Because few algae species exhibit the red body characteristic of N. oceanica, its role in the organism’s cellular life cycle has remained largely unknown. This pigmented organelle initially forms adjacent to the cell’s food-producing plastid before ultimately being released outside the cell wall during autosporangial division. A parent N. oceanica alga divides into daughter cells via spore production in a process known as autospore release. Scientists’ interest in Eustigmatophyte algae is two-fold: its rapid growth and its ability to partition up to half its mass into valuable lipids. These hallmark features indicate the algae’s potential for use in biofuel applications. A range of reference genomes have been published as interest has grown in the potential application of gene editing tools to Eustigmatophyte algae toward biofuel applications. Once thought rare, Eustigmatophytes have been found in a range of freshwater, marine, and terrestrial systems. Two genera of Eustigmatophyte, Nannochloropsis and Microchloropsis, have been established as model systems. In both groups, cells are solitary, non-motile, and round, with diameters on the order of 2 to 4 microns. Species belonging to both genera reproduce on a diurnal basis via asexual fission, growing during the day and splitting each night. The presence of a red-orange globule outside the cell’s chloroplast during the daytime part of the life cycle is an identifying feature of Eustigmatophytes. Although the globule has been widely reported throughout the group, this work provides the first account of its formation and biological function. Imaging techniques included ultra-performance liquid chromatography coupled with high-resolution mass spectrometry, various laser and electron microscopy methods, and Fourier transform infrared spectroscopy. The intriguing autofluorescent, globular nature of the red body — with its distinct compartmentalization and differentiation — led researchers to define it as a membrane-bound organelle. Further study of the red body’s contents led researchers to hypothesize that it could be a delivery vessel for molecules used in cell wall construction. During the daytime portion of the life cycle, N. oceanica cells grow rapidly. At night, each cell divides into multiple daughter autospores. With the production of multiple daughter cells instead of just one, significantly more cell wall material is needed to fully encapsulate each new cell. The red body aids the encapsulation process, researchers believe, by making available large amounts of material to make a specific part of the cell wall known as algaenan. Infrared spectroscopy analyses back up this hypothesis, revealing that red bodies discarded after autospore generation contain a range of precursor and intermediate products needed for cell wall formation, in addition to some fully polymerized algaenan matter. Researchers see N. oceanica as a model organism for understanding the biosynthesis of “chemically recalcitrant lipidic biopolymers” via plastid-derived fatty molecules that must be transported through an aqueous inner-cell environment to the cell wall. Similar molecules are ubiquitous throughout plant lineages because they play a key role in plant physiology by controlling the movement of water within and around plant bodies and the cells that constitute them. Until now, many details related to the transport of these molecules within the cell, and their final polymerization process during cell wall construction, remained unknown. As a result, this work contributes new insight into the biological link between molecular and large-scale processes at the cellular level. | |
| 11/28/2023 | Spatiotemporal Insights into Cellulose Hydrolysis | Berkeley Synchrotron Infrared Structural Biology Imaging Program | Synchrotron Infrared Hyperspectral Imaging | Advanced Light Source | Chemical and Elemental Information | With the recent introduction of closed microfluidic systems into synchrotron-based radiation Fourier-transform infrared (SR-FTIR) spectroscopy techniques, researchers have new options for non-destructively probing sub-cellular systems at high spatial and temporal resolutions. FTIR provides insights into chemical bonds that typify systems and changing patterns of chemical bonds; it has the potential to elucidate the biochemical dynamics of living systems in response to changing stimuli and other environmental cues. This study took microfluidic methods one step further by employing a novel open-microfluidic device, which enables on-demand access to the sample environment during infrared analysis while autonomously maintaining the ultra-thin liquid films required for efficient infrared analysis. The results serve as a key proof-of-concept for the technique, with the potential to enable a new generation of scientific discoveries—especially for researchers seeking to characterize the physical and chemical properties of biological systems in situ and in real time. The study also sheds new light on the spatiotemporal dynamics of enzymatic hydrolysis in cellulose, an important step toward harnessing this important biofeedstock toward a circular bioeconomy. | The structure of cellulose is predominated by highly-ordered rope-like structures known as fibrils, making this biopolymer an intriguing feedstock for manufactured nanomaterials with the potential for impressive mechanical properties. Additionally, cellulose depolymerization yields glucose, which can be used to produce biochemicals and biofuels. But the intra- and intermolecular hydrogen bonds that give cellulose fibrils their impressive mechanical properties also reduce the accessibility of enzyme interaction sites, limiting the efficiency of the enzyme-driven hydrolysis steps needed to process cellulose biomass into usable products. Research from the University of California-Davis and the Berkeley Synchrotron Infrared Structure Biology Imaging Program (BSISB) at Lawrence Berkeley National Laboratory builds on prior research to better understand the dynamics of enzyme-driven cellulose hydrolysis by introducing a breakthrough open-microfluidic device. The device enables real-time, in situ characterization of these important biochemical processes, which were previously thought impossible to perform, when coupled with non-destructive, synchrotron-generated infrared light. The first-of-its-kind microfluidic device enables tailoring of the sample environment, revealing how recalcitrant cellulose fibrils respond to cellulases. The technology can easily be extended to other biomolecular systems of interest to provide insight through real-time investigations at high spatial resolution. | Many studies and scientific techniques have provided important insight into enzymatic interactions with cellulosic biomass. Even so, the role of the cellulose supramolecular structure (i.e., its fibrils and hydrogen bonds) in the efficiency of such enzymatic interactions remains unclear. The best way to gain deep insight into these processes is through in situ, time- and spatially-resolved characterization techniques. Hence, researchers in this study coupled a novel open-channel, high-density capillary microfluidic device with existing SR-FTIR techniques. Until now, the enzymatic hydrolysis of cellulose was a challenging biochemical process to study, in part due to limitations in researchers’ abilities to carefully control and observe the sample environment (e.g., reactive fluids or relative humidity). The new technique overcomes these challenges through several approaches. First, the exceptional brightness of the synchrotron infrared light source supports real-time data capture. Second, the open nature of the microfluidic device enables on-demand access to the sample environment with autonomous and precise control of relative humidity, even as more reactive fluid is added. As a result, the approach supports a sustainable technique for detecting key spectral signals as the biochemical reaction of interest proceeds within the microfluidic sample environment. This approach could be extended across a range of systems and sample types to reveal insights previously thought inaccessible to science. In this work, the time-resolved depletion of algal cellulose by a purified cellulose-degrading enzyme was tracked by monitoring a set of absorption peak intensities associated with the glycosidic bond—the covalent bond that joins glucose monomers within the cellulose structure. The results demonstrated high spatial heterogeneity of the enzymatic process within the sample, found to be consistent with the notion that fibrillar regions containing large amounts of intra- and intermolecular hydrogen bonding are able to resist enzymatic processes. This insight can help researchers devise new approaches for cellulose biomass processing, with the ultimate goal of developing a range of manufacturing applications toward a more sustainable and circular bioeconomy. Advances in closed and open high-density microfluidic technologies developed through BSISB-led research and development initiatives can directly benefit BSISB users who require controllable sample environments or reaction conditions, as well as in situ monitoring at high spatial resolution. | |
| 04/12/2024 | Nitroplast: Symbiont-Turned-Organelle Fixes Nitrogen for Alga | National Center for X-Ray Tomography | Soft X-ray Tomography | Advanced Light Source | Cell and Tissue Structure | The marine alga B. bigelowii is the first known eukaryote to pull nitrogen from the air. As reported in a Science perspective article, the “new data support the claim that nitrogen fixation is no longer an exclusive prokaryotic function and that eukaryotes can fix molecular nitrogen through the nitroplast. The nitroplast represents a textbook case of a eukaryotic organelle that complements the energy, carbon, and nitrogen needs of the algal host and is another example of how ecology is the theater where evolution takes place.” The discovery is of great interest for understanding organelle genesis and for efforts to engineer agricultural plants with built-in nitrogen-fixing capabilities. | Eukaryotic cells differ notably from prokaryotes in that they contain membrane-bound organelles with specific functions. A few of these organelles, such as mitochondria and chloroplasts, started out as endosymbiotic bacteria but evolved to become fully integrated into the host cell. An international team of scientists from the University of California at Santa Cruz, Lawrence Berkeley National Laboratory, and other institutions now report close integration of the nitrogen-fixing cyanobacterial endosymbiont Candidatus Atelocyanobacterium thalassa into the architecture and function of its unicellular marine algae host Braarudosphaera bigelowii. Proteomic and X-ray tomographic evidence reveal that this close integration is more characteristic of an organelle than an endosymbiont. The findings suggest that Candidatus A. thalassa evolved from a symbiont to a eukaryotic organelle for nitrogen fixation—named the nitroplast—thereby expanding to eukaryotes a function that was thought to be exclusively carried out by prokaryotic cells. | Evidence distinguishing the nitroplast nitrogen-fixing organelle from an endosymbiont comes from proteomics and soft x-ray tomography (SXT) conducted at the National Center for X-ray Tomography at the Advanced Light Source. SXT shows that division of the nitroplast organelle is coordinated with that of the other organelles and omics studies indicate that half of the nitroplast proteins were imported from the B. bigelowii host genome nucleus to fill in gaps in its metabolic pathways. Rather than symbiont genomes migrating into the host’s nuclear genome, the host genome appears to support the symbiont. This suggests that the nitroplasts result from primary endosymbiosis—the process by which a symbiont is engulfed by a host organism and evolves beyond symbiosis to become part of the host cells. This evolutionary process is analogous to what occurred with mitochondria and chloroplasts. The discovery raises the prospect of engineering plants that can fix nitrogen directly from the air, reducing the need for synthetic nitrogen fertilizers, the production of which generates enormous amounts of carbon dioxide. | |
| 11/22/2023 | Deuterium Labeling Reveals Structural Components of Plant Cell Walls | Center for Structural Molecular Biology | Small-Angle Neutron Scattering, Synchrotron Infrared Hyperspectral Imaging | Spallation Neutron Source/High Flux Isotope Reactor | Molecular Structure | Deuterium labeling increases the difference in neutron scattering length density between carbohydrate and lignin plant polymers, allowing deconvolution of the structural features of different component biopolymers without altering native cell wall structure. This approach will improve understanding of how cellulose, hemicellulose, and lignin interact during plant cell wall assembly as well as the structural changes that occur during biomass deconstruction for biofuels production. | Plant cell walls have a complex structure consisting primarily of cellulose, hemicellulose, and lignin. These polymers come together to form an intricate laminate that confers an architecture advantageous for producing a strong and durable material. However, the interactions between the polymers and their distributions in cell walls are not well understood. Plants grown in a mixture of water and heavy water (deuterium oxide; D2O) differentially incorporate deuterium into their cell wall polymers. Fourier transform infrared spectroscopy (FTIR) revealed that carbohydrate polymers incorporate more deuterium than lignin. Using this differential, structural characterization could be performed using small-angle neutron scattering (SANS), which demonstrated that cellulose and lignin fractions in cell walls can be spatially deconvoluted. This approach can help improve understanding of polymer organization in plant cell walls and the changes that occur during their deconstruction to produce biofuels. | Brassica Oleracea (kale), a relative of bioenergy crops B. napus (rapeseed), and B. carinata (camelina) and a C3 plant (e.g., like poplar and most trees), was grown in a mixture of water and D2O. Incorporation of deuterium in the different plant polymers was determined using FTIR. Contrast variation small-angle neutron scattering (CV-SANS) was then used to structurally characterize the plant biomass. FTIR results showed higher deuterium incorporation in the carbohydrate fraction (~41%) compared to lignin (21%). CV-SANS showed it was possible to separate the scattering signals of the different polymers, which is not possible using unlabeled plant cell walls. Lignin dominated the scattering intensity at longer spatial scales while cellulose was observed at shorter length scales. | |
| 10/23/2023 | Lignin−Pectin Complexation Implicated in Biomass Recalcitrance | Center for Structural Molecular Biology | Small-Angle Neutron Scattering, Solution X-ray Scattering, Synchrotron Infrared Hyperspectral Imaging | Spallation Neutron Source/High Flux Isotope Reactor | Molecular Structure | This work shows that interactions between pectin and lignin may be a previously unidentified contributor to LCCs in plant cell walls. Engineering plants to reduce these interactions could decrease biomass recalcitrance for biofuel production. | Lignin–carbohydrate complexes (LCCs) form through interactions of lignin with plant cell wall polysaccharides and are thought to be a significant source of biomass recalcitrance. Understanding LCCs and their effects on lignin morphology is needed to improve biomass conversion to biofuels and bioproducts. The role of pectin plant cell wall recalcitrance was investigated and showed strong evidence that pectin changes how lignin distributes in plant cell walls during thermochemical pretreatment. Additional analysis of a model composite composed of pectin and lignin showed that pectin and lignin form a highly interconnected polymer network and evidence of an ester bond between the polymers. Overall, this study provides new insights into the relationship between primary and secondary cell wall polymers during cell wall synthesis. It may help develop new approaches to modulate cell wall properties to improve biofuel and bioproduct production. | This work investigated LCCs formed between lignin and the pectin homogalacturonan (HG). Structural changes in HG-deficient transgenic switchgrass after hot water pretreatment were compared to wildtype plants using small-angle neutron scattering (SANS). SANS showed ~2.2-fold more lignin aggregates in the transgenic biomass compared to wildtype. This suggests that decreased pectin resulted in greater lignin redistribution to form aggregates and that interactions between lignin and HG restrict lignin mobility in plant cell walls. To better understand the types of interactions between lignin and pectin, a model composite was prepared by polymerizing either protiated or partially deuterated coniferyl alcohol to form dehydrogenation polymer (DHP) in the presence of HG. Small-angle X-ray scattering showed that the DHP and HG form a highly interconnected network structure that is not observed in a physical mixture of the individual polymers. Contrast matching SANS revealed the structural characteristics of DHP and HG in the composite and showed that the HG forms a swollen interconnected polymer network interspersed with DHP particles that are composed of solvent-accessible DHP polymers. Fourier transform infrared spectroscopy showed a unique ester absorption band in the DHP/HG composites. Solid-state nuclear magnetic resonance (NMR) analysis also supports interactions between DHP and HG. | |
| 03/27/2023 | Computational Tool Optimizes SANS Experiments | Center for Structural Molecular Biology | Small-Angle Neutron Scattering | Spallation Neutron Source/High Flux Isotope Reactor | Molecular Structure | SCOMAP-XD calculates the contrast-match points and Q-dependent contrast for experimental optimization of biomacromolecular neutron scattering experiments. Previous methods have only used random assignment or bulk solute contrast effects for determining contrast match points. This approach significantly advances understanding of SANS contrast and enables further investigation into Q-dependent contrast effects. The approach is readily applicable to a large variety of systems, including polymer and carbohydrate systems like cellulose. | Structural characterization of individual proteins, lipids, and nucleic acids within biomacromolecular complexes can be achieved using small-angle neutron scattering (SANS) and isotopic labeling. To optimize design of SANS experiments, this study developed a contrast matching calculation workflow, called Scattering Contrast Match Points with Explicit-atom Deuteration (SCOMAP-XD), using atomic simulations with explicit-atom deuteration. Contrast matching is central to SANS because it enables different components of a biological system to be investigated individually. Biological macromolecules are typically dissolved in water containing both hydrogen and deuterium (i.e., heavy water; deuterated water; D2O), which produce different scattering signals. By manipulating the ratios of hydrogen to deuterium in the solution until it progressively reaches match points for different biomolecules, or by manipulating ratios in a specific molecule itself, the system’s different macromolecular components can be observed one at a time. SCOMAP-XD is a computational tool that explicitly deuterates biomolecular three-dimensional structures to determine the overall contrast match point and the Q-dependent contrast-match points of biomacromolecules. In addition, SCOMAP-XD combines information from empirical models to understand deuterium incorporation at exchangeable and nonexchangeable hydrogen sites in molecules, due to hydrogen-deuterium exchange and the deuterium oxide concentrations in the cell culture media used for biomacromolecule production. | The SCOMAP-XD method incorporates empirical models for determining the incorporation of deuterium at non-exchangeable hydrogens from the deuterium oxide (D2O) concentration in the culture medium using mass spectrometry and for determining the probability for hydrogen-deuterium exchange at exchangeable hydrogens from nuclear magnetic resonance hydrogen-deuterium exchange data. Using a 3D structure, hydrogen sites for deuterium incorporation can be selected and scattering profiles can be calculated using the explicit solvent calculator, SASSENA. From the calculated structures, the contrast match points can be calculated within 2.5% accuracy. In addition, scattering-vector-dependent contrast effects were observed at high Q in several systems, which are indicative of specific contributions due to deuterium distribution in the sample. | |
| 01/24/2023 | SANS Reveals Role of Disordered Protein Domain | Center for Structural Molecular Biology | Small-Angle Neutron Scattering | Spallation Neutron Source/High Flux Isotope Reactor | Molecular Structure | This study shows for the first time that a disordered region of c-Src kinase modulates the structure of the neighboring folded domain. The observation may have important implications for allosteric interactions with binding partners. This type of structural information is difficult if not impossible to obtain with other structural characterization techniques using X-rays and electrons. The segmental labeling approach can be broadly applied to study functionally important disordered regions in other multidomain proteins involved in cell signaling and other biological processes in biological and environmentally relevant systems. | Researchers at the Center for Structural Molecular Biology and the Center for Biophysics at Oak Ridge National Laboratory developed a segmental labeling procedure to study interactions between an intrinsically disordered N-terminal protein domain and folded domains in the multidomain protein c-Src kinase. The procedure, called domain-selective deuterium isotopic labeling or segmental labeling, was combined with small-angle neutron scattering (SANS). Together, these techniques enable functionally important disordered regions to be studied in multidomain proteins, providing structural insights that are difficult if not impossible to obtain with X-ray-based or electron-based structural characterization techniques. | The protein c-Src kinase is a multidomain non-receptor tyrosine kinase associated with many types of cancer. Although the structural properties of the protein’s folded catalytic and regulatory domains (SH3-SH2) have been extensively characterized, less is known about the N-terminal disordered region (SH4UD). Protiated SH4UD was enzymatically ligated to deuterated SH3-SH2 domains to synthesize a single polypeptide chain of (SH4UD)H-(SH3-SH2)D. Contrast variation SANS showed that in the presence of SH4UD, the radius of gyration (Rg) of SH3-SH2 increases, indicating that it has a more extended conformation. Hamiltonian replica exchange molecular dynamics simulations provide a detailed molecular description of the structural changes in SH4UD-SH3-SH2. The simulations showed that the regulatory loops of SH3 undergo significant conformational changes in the presence of SH4UD while SH2 remains largely unchanged. | |
| 11/27/2023 | SAXS Elucidates a Microbial Bioenergetic Pathway | Berkeley Synchrotron Infrared Structural Biology Imaging Program | Solution X-ray Scattering | Advanced Light Source | Molecular Structure | By following the mechanics and conformations of EtfABCX, the researchers were able to shed light on this novel biological mechanism where energetic electrons are channeled over relatively large distances. Understanding these novel bioenergetic systems will help build microbial energy flux models and aid in the annotation of newly discovered genes. | Electron bifurcation, a class of chemical reaction found only in biology, liberates two electrons from one donor and channels them to two separate electron acceptors. Analogous to a mechanical pulley system in which one weight rises as the other falls, one electron is elevated in energy at the expense of lowering the energy of the second. The higher-energy electron can then help perform high-energy work, such as bond-breaking or bond-making, as necessary for the organism’s survival. Now, a team of researchers from the Lawrence Berkeley National Laboratory and other institutions has used small-angle x-ray scattering (SAXS) at the Advanced Light Source (ALS) to understand an important protein involved in this bioenergetic pathway—the EtfABCX protein from the anaerobic bacterium Thermotoga maritima. In experiments performed at ALS Beamline 12.3.1 (SIBYLS), the team studied the mechanism of electron channeling in the bifurcation process. SAXS, a solution-based technique that does not restrain motion, enabled researchers to elucidate the steps, triggers, and coordinated conformational changes of EtfABCX. The results suggest that the protein couples electron transfer to conformational change. | ||
| 10/16/2023 | Computer-Aided Protein Design for New Biomaterials | Structurally Integrated Biology for the Life Sciences | Solution X-ray Scattering | Advanced Light Source | Molecular Structure | This study’s computational approach enables highly accurate design of protein crystals. The designed crystals, which are encoded with both structural and assembly information in their primary sequences, provide a powerful platform for biological materials engineering. | Protein crystallization plays a central role in structural biology research, yet the crystallization process remains poorly understood and highly empirical. Crystal contacts, lattice packing arrangements, and space group preferences are largely unpredictable. Programming protein crystallization through precisely engineered side-chain–side-chain interactions across protein–protein interfaces is therefore an outstanding challenge. Efforts to understand protein structure and manipulate function typically begin with crystallization followed by X-ray diffraction to determine the molecular arrangement of atoms. Computational protein design now enables scientists to explore ways to use proteins to create completely new materials with precisely tuned characteristics such as lattice dimension and pore size. Using a computer-based approach, researchers designed porous protein crystals that were revealed to be stable, tunable, and atomically accurate using X-ray scattering and diffraction at the Advanced Light Source (ALS). | This study developed a general computational approach for designing three-dimensional protein crystals with prespecified lattice architectures at atomic accuracy that hierarchically constrains the overall number of degrees of freedom of the system. Researchers designed three pairs of oligomers that can be individually purified and spontaneously self-assemble into >100 µm three-dimensional crystals upon mixing. The crystal structures are nearly identical to the computational design models, closely corresponding in both overall architecture and the specific protein–protein interactions. The crystal unit cell dimensions can be systematically redesigned while retaining space group symmetry and overall architecture, and the crystals are extremely porous and highly stable. | |
| 12/22/2023 | Sulfur Exchanged in Peat Moss-Cyanobacteria Mutualism | Structural Molecular Biology Resource | X-ray Fluorescence Imaging | Stanford Synchrotron Radiation Lightsource | Chemical and Elemental Information | Whether peatland ecosystems continue to serve as net carbon sinks or become carbon sources as a result of climate change will, in part, depend on how Sphagnum-microbe interactions respond to various climate factors. The current understanding of the Sphagnum–cyanobacterium symbiosis is that carbon-rich carbohydrates are exchanged for nitrogen-rich molecules, but this conceptual model appears to be overly simplistic and possibly missing a key role for sulfur. This study’s observations emphasize the potential importance of sulfur metabolite production by Sphagnum for facilitating symbiotic association with nitrogen-fixers. Future studies could enable a better understanding of how Sphagnum-driven sulfur dynamics influence associated carbon and nitrogen inputs to peatland ecosystems. | Peatland ecosystems greatly impact terrestrial carbon and nitrogen processes, occupying 3% of the Earth’s land surface and storing approximately 25% of terrestrial carbon as recalcitrant organic matter (OM). Sphagnum, a key peat moss genus and producer of recalcitrant OM, is responsible for much of the primary production in these ecosystems. Its growth and productivity depend on symbiotic associations it forms with various microbes, including cyanobacteria.  Fluorescence images showing sulfur speciation in Sphagnum angustifolium leaves colonized by cyanobacteria. [Reprinted with permission from Weston et al. 2023. New Phytologist, DOI:10.1111/nph.19476. Copyright 2023 John Wiley and Sons.] The mutualistic peat moss-cyanobacteria relationship appeared to at least temporarily increase oxidized sulfur compounds within Sphagnum tissues that subsequently undergo anaerobic decomposition in saturated peat soils. Additional studies are needed to determine the role of sulfur exchange in driving ecosystem-scale sulfur, nitrogen, and carbon dynamics. | Sphagnum angustifolium (originally collected in the field) were colonized by Nostoc muscorum by first adding N. muscorum to a 12-well plate. S. angustifolium gametophytes were placed in a fitting cell culture insert and then placed in a Nostoc-filled well for colonization. Experiments ended after 14 days and each leaf was examined under an epifluorescence microscope equipped with a green (to show plant material) and red (to show cyanobacteria) excitation filter. Percent colonization was quantified as the percent of S. angustifolium hyaline cells occupied by N. muscorum. Samples were prepared for X-ray analysis by rinsing each leaf (or in some cases leaf clusters) in deionized water, then gently placing each sample onto a microscope slide and covering with sulfur-free tape prior to shipping to the Stanford Synchrotron Radiation Lightsource (SSRL). At SSRL, the sulfur-free tape was gently removed with the leaf adhered to the tape, and the tape was loaded onto a sample holder for analysis at beamline 14-3. For each sample, multiple-energy (ME) maps across the sulfur K-edge were collected over the entire life (or cluster) to map the spatial distribution of sulfur species. Sulfur XANES spectra were collected across the samples, with spots determined by on-the-fly analyses of ME maps. The data showed the presence of reduced organic sulfur and oxidized sulfonate- and sulfate-containing compounds. The abundance of these sulfur species changed with percent colonization, where an increase in colonization by N. muscorum resulted in an enrichment of sulfur and changes in speciation, with increases in sulfate relative to reduced sulfur and sulfonate. At the scale of individual hyaline cells, colonized cells exhibited localized enrichment of reduced sulfur surrounded by diffuse sulfonate, similar to observations of N. muscorum colonies alone. | |
| 07/21/2023 | Developing an <i>E.coli</i> Platform for Hydrogenase Production | Structural Biology Center | X-ray Macromolecular Crystallography | Advanced Photon Source | Molecular Structure | The genetic constructs created in this study, together with improved growth and purification procedures, provide a promising platform for further studies toward producing fully active and oxygen-tolerant recombinant Ni-Fe-Se hydrogenase resembling the native D. baculatum enzyme. | Hydrogenases are metalloenzymes, found in microbes, which efficiently convert protons and electrons to molecular hydrogen. Industrial production of hydrogen, a potential fossil fuel alternative, requires intensive use of chemicals, toxic metals, and energy. Developing an industrial process that exploits the unique enzymatic activity of microbial hydrogenases could advance the process of biohydrogen evolution and green energy production. Researchers from the University of Gdansk and Argonne National Laboratory used site-directed mutagenesis of a bacterial hydrogenase operon—a cluster of genes that work in concert to produce the enzyme—and molecular imaging to create a functional, recombinant operon for rapid and robust hydrogenase production in the industrial powerhouse E. coli. Using such strains alleviates the challenging growth constraints of much slower growing and difficult to cultivate native sources of hydrogenase—such as the anaerobic, sulfate-reducing Desulfomicrobium baculatum used in this study. The resulting recombinant hydrogenase varies from the native enzyme in that its active site is composed of nickel and iron (Ni-Fe), whereas the native enzyme has a Ni-Fe-Selenium (Ni-Fe-Se) core. The Ni-Fe hydrogenase displays only a fraction of the native enzyme’s activity, and its properties still need to be evaluated, but the study demonstrates a promising strategy for cloning, expressing, and purifying catalytically active hydrogenase derived from D. baculatum in E. coli. | The native Ni-Fe-Se hydrogenase from D. baculatum was targeted in this study for its natural oxygen tolerance and high hydrogen evolution activity. The native hydrogenase operon includes two structural hydrogenase genes, coding for large and small subunits, and an additional gene, encoding a protease essential for proper enzyme maturation. Recombinant expression of these exact genes in E. coli, however, produces inactive enzymes. Therefore, researchers converted the native Ni-Fe-Se hydrogenase to an Ni-Fe type hydrogenase using site-directed mutagenesis. The four resulting recombinant hydrogenase variants demonstrated limited ability to produce hydrogen both in vitro and in vivo. The aim of this study was to overcome bottlenecks in gene expression, protein biosynthesis, maturation, and ligand loading for simple, rapid, and cost-effective delivery of recombinant Ni-Fe hydrogenase using commonly available E. coli strains. Although the enzymes produced in this study do not yet rise to the level of the native enzyme, the demonstrated platform for recombinant hydrogenase production shows promise. Improvements could likely be achieved by selective cysteine to selenocysteine substitution within the active site of the Ni-Fe variant. | |
| 08/07/2023 | Structural Analysis of Neglected Anaerobic Fungal Enzymes | eBERlight, Structural Biology Center | X-ray Macromolecular Crystallography | Advanced Photon Source | Molecular Structure | The atomic-resolution structure and detailed biochemical characterization of CelD add to a growing understanding of how anaerobic fungal cellulosomes rapidly degrade biomass. Mining anaerobic fungal genomes for better lignocellulolytic enzymes than traditional aerobic microbes may yield higher performing enzymes for industrial biomass valorization applications. Characterizing the atomic-resolution structure and kinetic properties of the P. finnis CelD GH5 endoglucanase provides additional insight towards gaining biochemical understanding of anaerobic fungal enzyme systems. | Microbial degradation of lignocellulose is a fundamental biological process crucial to nutrient cycling in nature. The process turns over the equivalent energy of an estimated 640 billion barrels of oil per year. Various microbial enzymes work synergistically to break down hemicellulose, cellulose, and lignin in plant cell walls, with carbohydrate-active enzymes (CAZymes) of the glycoside hydrolase (GH) family largely responsible for cellulose saccharification. Anaerobic fungi frequently demonstrate cellulolytic activity and hemicellulose degrading power that equals or exceeds the highest performing aerobic microbes and enzyme cocktails. The genomes of these prolific biomass degraders, often found in the guts of large herbivores, harbor a potential wealth of industrially relevant CAZymes. However, very few have been structurally or biochemically characterized, so very little is known about the characteristics that render the enzymes so efficient. Such knowledge gaps in the biochemistry of individual anaerobic fungal CAZymes themselves, and of fungal cellulosomes which comprise many CAZymes, present a challenge towards designing lignocellulolytic enzyme cocktails that leverage the degradative machinery of anaerobic fungi into useful biotechnologies. Scientists from Argonne National Laboratory, the University of California-Santa Barbara, and colleagues from three other institutions used X-ray macromolecular crystallography and additional techniques to begin filling the knowledge gap by determining the structure and kinetic rate parameters for a GH family 5 subfamily 4 enzyme, called CelD, from the unique anaerobic fungus Piromyces finnis isolated from the guts of large herbivorous mammals. Notably, CelD’s kinetics do not change with domain fusion, which suggests it is highly modular. | The study presents the crystal structure of a CelD GH5 catalytic domain in complex with cellotriose, a carbohydrate substrate. CelD is a cellulase that acts strictly as an endo-β-1,4-glucanase with confirmed activity against carboxymethylcellulose, mixed linkage glucan, and xyloglucan. Observing structural changes during cellotriose substrate binding by the enzyme and linking these structural transformations with the enzyme’s kinetics provided insight into CelD’s catalytic mechanism. The structure presents a platform for rational engineering of this enzyme for higher affinity, thermostability, or activity criteria. The kinetic data indicate that the CelD domains are highly modular and can likely be augmented to be functionalized with other domains that may act synergistically. | |
| 06/12/2023 | Visualizing an Ancient Carbon Cycle Pathway Mechanism | Structural Molecular Biology Resource | X-ray Absorption and Emission Spectroscopy | Stanford Synchrotron Radiation Lightsource | Chemical and Elemental Information | This work details the remaining undescribed pieces of an ancient and critically important carbon fixation pathway using XAS and provides a new conceptual framework to bridge two previously competing mechanistic models of acetyl-CoA synthase (ACS). | The Wood-Ljungdahl pathway (WLP) is one of nature’s most elegant methods for chemically transforming carbon. The pathway is ancient, present in the last universal common ancestor to life on earth, and the only carbon fixation pathway known to produce more energy (net ATP) than it consumes. The WLP scrubs CO2 from the atmosphere, converting the carbon to energy and cellular building material. The process occurs within a large multi-unit protein complex comprising carbon monoxide dehydrogenase (CODH), which turns CO2 into CO, and acetyl-CoA synthase (ACS), which combines CO and a CH3– group donated by a corrinoid iron-sulfur protein (CFeSP) to produce acetate in the form of acetyl-CoA from the original CO2 molecule. Globally, anaerobes produce 1013 kg of acetate annually via the WLP. The pathway represents an ideal system upon which to develop biomimetic catalysts for atmospheric CO2 capture, but its underlying mechanism must first be understood. Unique nickel (Ni)-based and iron-based active sites lie at the hearts of CODH and ACS where acetate and acetyl-CoA form through a series of Ni-based organometallic intermediates. The exact mechanism of acetate synthesis from CO and CH3 has been intensely studied for several decades, with two proposed mechanisms at the center of debate. Both propose that CO and CH3 reactants bind to a single nickel site (termed Nip), but each proposes a different oxidation state and electronic configuration for the reactant-bound Nip. In the paramagnetic mechanism, CO and CH3 react with Nip to form a series of paramagnetic Ni(I) and Ni(III) intermediates; in the other mechanism, catalytically active Nip species are formally diamagnetic Ni(0) and Ni(II). In a collaborative study supported in part by the Structural Molecular Biology Resource at the SLAC National Accelerator Laboratory, researchers solved the geometric and electronic structures of the remaining uncharacterized organometallic intermediates of ACS using X-ray absorption spectroscopy (XAS) and other techniques. The structures revealed that ACS binds CO, CH3, and acetate through direct Ni-C bond formation, and that the Ni-CH3 intermediate is paramagnetic. In addition, the paramagnetic intermediates transformed under experimental conditions into their diamagnetic analogs and these diamagnetic species appeared to be recruited into the catalytic cycle as well. To explain the existence of both catalytically relevant paramagnetic Ni(III) and diamagnetic Ni(II) species, the researchers proposed a novel electrochemical coupling mechanism. Under this mechanistic framework, Nip(I) (or Nip(I)-CO) reacts with methyl-Co(III) to generate Nip(III)-methyl (or Nip(III)-acetyl), which are reducible to their Nip(II) counterparts via interaction with excess reductant. These reduced intermediates are then recruited back into active catalysis through binding of the next substrate. | ||
| 06/06/2023 | Automation and High-Throughput Screening Speeds Lipid Nanoparticle Development | Structurally Integrated Biology for the Life Sciences | Cryo-Electron Microscopy, Solution X-ray Scattering | Advanced Light Source | Molecular Structure | This method can be applied to rapidly characterize and optimize countless LNP formulations for different applications. LNP morphology is recognized as a critical parameter governing LNP bioactivity, but structural analysis is typically not assessed due to limited accessibility and resource requirements. The high-throughput LNP formulation and screening workflow developed in this study enables researchers to produce and characterize LNPs at record speed. The study also includes the first-ever demonstration of how LNP structure correlates with the activity of its contents. In addition, unlike other forms of X-ray diffraction on biological materials, SAXS samples do not require freezing or crystallization which can change LNP structure. SAXS also enables snapshots of LNPs at a specified timepoints to determine structural longevity. |  Diverse research areas are exploring potential applications of lipid nanoparticles (LNPs). Although pharmaceutical uses currently dominate the LNP space, as represented here by the percentage of all LNP-related scientific publications from 2000 to 2020, many other top research areas are gaining ground. BER-relevant research areas include biochemistry, biochemical methods, genetics, agriculture, and toxicology. [Reprinted under a Creative Commons license (CC BY 4.0) from Tenchov et al. 2021. DOI:10.1021/acsnano.1c04996] LNPs are nanoscale spherical pouches composed of a lipid membrane encapsulating various cargo for delivery to cells. Most prominently, LNPs serve as delivery systems for therapeutic agents, including mRNA in two SARS-CoV-2 vaccines. The LNPs protect the mRNA or other cargo from enzymatic degradation inside the body and prevent it from adversely affecting non-target tissues and systems. Beyond therapeutics, LNP applications have been extended to medical imaging, cosmetics, nutrition, and agriculture. They are also being studied as delivery systems for agrochemicals; as model membrane systems; and as nanoscale chemical reactors applied in nanotechnology and nanobiotechnology. The success of LNPs in medicine motivates further fundamental and applied LNP research in the fields of environmental science, such as metal detoxification and the manipulation of microbial communities, and materials science, such as the controlled synthesis of metal nanoparticles. This study relied on high quality and high quantity SAXS data generated at the Structurally Integrated Biology for the Life Sciences resource, as well as cryogenic electron microscopy to verify the SAXS data. | LNPs are comprised of four major components, allowing them to be customized in countless permutations. These components include ionizable lipids, helper phospholipids, cholesterol, and polyethylene glycol-lipids (PEG-lipids). Capitalizing on this flexibility, researchers used an automated process to rapidly generate a library of LNPs containing potentially therapeutic antisense oligonucleotides (ASO). Structural characterization of the resulting ASO-LNPs was performed using high-throughput SAXS backed up by cryogenic electron microscopy (cryo-EM). Associating ASO-LNP SAXS signals to their respective cryo-EM features enabled identification of LNP structural parameters that can be analyzed directly from SAXS data. The structural information was then combined with the results of a high-throughput in vitro efficacy assay in mouse cortical neurons to determine any relationship between ASO-LNP structure and cellular efficacy. The method yielded a structure–activity relationship for LNPs in a high-throughput setup to help identify chemical compositions that produce optimal LNP structures for maximized efficacy. | |
| 06/30/2023 | Persistence and Mobility of Iron-Rich Colloids Facilitate Element Transport and Cycling | Stanford-SLAC Cryo-EM Center | Cryo-Electron Microscopy, X-ray Absorption and Emission Spectroscopy | Environmental Molecular Sciences Laboratory, Stanford Synchrotron Radiation Lightsource | Chemical and Elemental Information | The ability of ferrihydrite-based colloids to withstand anoxic conditions rich in dissolved Fe(II) highlights the extent to which silicon and organic matter coatings can protect Fe(III) from reductive dissolution. This passivating (i.e., corrosion deterring) feature may also explain the existence of Fe(II) and sulfur within the colloidal structure. Ultimately, the persistence of the colloids suggests they may transport throughout anoxic zones and reach oxic surface waters. The findings shed light on the composition and dynamics of natural Fe-rich colloids in floodplain systems, with implications to elemental transport and cycling. | Subsurface interfaces are ubiquitous in floodplain environments and sustain a multitude of biogeochemical processes, including the formation and release of reactive, mobile colloids. Colloids are known vectors of micronutrient, contaminant, and organic matter transport and are suspected to be important export agents from floodplains to stream water. Iron (Fe)-rich mobile colloids play vital yet poorly understood roles in the biogeochemical cycling of Fe in groundwater by influencing organic matter (OM) preservation and fluxes of Fe, OM, and essential micronutrients. Researchers from the SLAC National Accelerator Laboratory, the Environmental Molecular Sciences Laboratory (EMSL), and other institutions detected Fe-rich colloids in anoxic groundwater of a redox-active floodplain near Slate River, Colo, where the colloids accounted for up to 72% of aqueous Fe. They characterized the colloids using a wide array of techniques including cryo-transmission electron microscopy (cryo-TEM), TEM-energy dispersive spectroscopy (TEM-EDS), Mössbauer spectroscopy, and Fe-extended X-ray absorption fine structure (Fe-EXAFS). The colloids comprise mixed-phase assemblages of ferrihydrite nanoparticles coated with silicon and enmeshed in organic matter. Both Fe(II) and Fe(III) co-existed in the colloids under anoxic conditions, illustrating the passivating (i.e., corrosion resistant) effects of the silicon and organic matter matrix against redox-triggered transformations. | Researchers used a combination of advanced characterization techniques to decipher the composition of Fe-rich colloids at a floodplain field site near Slate River, Colo. Cascade filtering revealed the presence of Fe-rich colloids in the riparian anoxic soil water and their abundance and composition varying with season. Cryo-EM and TEM-EDS imaging showed monodispersed and nanoassemblages of spherical colloids in the 10-50 nm range containing Fe, oxygen, silicon, carbon, and calcium. TEM selected-area electron diffraction analysis and Mössbauer spectroscopy indicated a poorly crystalline ferrihydrite-like phase. Fe-EXAFS further verified ferrihydrite and Fe(II)- and Fe(III)-organic matter interactions, as well as a small contribution from Fe-sulfur bonding. The analyses indicate that the colloids were primarily composed of nanosized ferrihydrite spheres stabilized by organic matter, silicon, and bridging cations such as calcium. The persistence of Fe(III)-rich colloids in primarily anoxic zones suggests that the silicon-organic matter coating serves as a passivating layer against reduction, but its efficiency likely depends on biogeochemical and hydrological conditions. | |
| 05/05/2023 | Saprotrophic Fungus Weathers Potassium from Minerals in Carbon-Limited Environment | Structural Molecular Biology Resource | X-ray Fluorescence Imaging | Environmental Molecular Sciences Laboratory, Stanford Synchrotron Radiation Lightsource | Chemical and Elemental Information | The observed mineral weathering patterns by fungal hyphae likely result from changing fungal metabolism with distance to a single carbon source. Distance affects the production of fungal exuded organic acids and drives surface mineral alteration and subsequently secondary mineral formation. This study details the transition of potassium from structurally bound, non-bioavailable potassium to readily accessible potassium pools, a key process for increasing local bioavailability of potassium in soils. | Potassium plays an important role in processes that contribute to overall plant health and growth and can mitigate drought effects in many species. Thus, finding sustainable pathways to increase bioavailability in the rhizosphere of this limiting environmental nutrient is critical as climate change continues to alter soil processes. Using micro-X-ray fluorescence (XRF) imaging combined with micro-X-ray absorption near edge structure (XANES) spectroscopy, researchers found that under nutrient-limited conditions the saprotrophic fungus Fusarium sp. DS 682 can indirectly weather potassium-rich minerals—specifically potassium feldspar and biotite—into tens-of-micrometer-scale clay particles that coat the mineral grains within 30 days. The distribution of clay coatings was associated with proximity to a carbon source, with more clay forming on potassium feldspar close to a carbon source but more clay forming on biotite at greater distances from the carbon source. | Researchers at the Environmental Molecular Science Laboratory (EMSL) developed synthetic soil habitats (SSHs) which replicate physical and chemical properties of soil within a reduced-complexity environment. A key aspect of SSHs is their amenability to multimodal imaging approaches. In this study, soil chemical properties were simulated using a mixture of potassium-rich minerals, such as potassium feldspar and micas, as well as kaolinite clay and quartz. After fungal growth, samples were lyophilized (i.e., freeze-dried) and the hyphae removed from SSH surfaces. The SSHs were cleaned with chloroform to remove any potassium sorbed to organic compounds on the SSH surface. This enabled the surface chemistry of the minerals to be determined without interfering signal from any leftover organics. The SSHs were then analyzed at the Stanford Synchrotron Radiation Lightsource (SSRL) X-ray microprobe beamline 14-3, which is well-suited to potassium chemical speciation imaging. | |
| 11/02/2022 | The Copper Key to More Efficient Biomass Breakdown | Center for Structural Molecular Biology | Neutron Macromolecular Crystallography | Spallation Neutron Source/High Flux Isotope Reactor | Molecular Structure | Understanding the mechanism of LPMOs could enable redesign and testing of different versions with improved efficiency for cellulosic ethanol production. | Nonfood, plant-based biofuels have potential as sustainable alternatives to fossil fuels, but the enzymes required for production are often too inefficient and costly to produce. However, new research shines a light on fungal enzymes called lytic polysaccharide monooxygenases (LPMOs) that could improve the economic viability of biofuels. LPMOs excel at breaking down organic matter, but their mechanism of action is unclear. Typical enzymes are made of carbon, nitrogen, hydrogen, and oxygen, but LPMOs also contain copper. Researchers had previously used neutron scattering at the Spallation Neutron Source and High Flux Isotope Reactor to show how LPMOs bind oxygen to copper to break down biomass. Going a step further, they’ve now used neutron protein crystallography to directly visualize protonation states in the initial steps of oxygen activation directly following active site copper reduction in LPMO9D from the fungus Neurospora crassa. The experiments reveal that the process is driven by an amino acid that donates protons to the oxygen molecule. | ||
| 03/28/2023 | Autonomous Hyperspectral Spatiochemical Imaging Delivers Faster Results and Improves Beamline Efficiency | Berkeley Synchrotron Infrared Structural Biology Imaging Program | Solution X-ray Scattering, Synchrotron Infrared Hyperspectral Imaging, X-ray Fluorescence Imaging | Advanced Light Source | Molecular Structure, Chemical and Elemental Information | Autonomous experimentation is an emerging area of research that has now been extended to infrared (IR) spatiochemical mapping of dynamic biological systems. Autonomous approaches have been difficult to apply to traditionally high-dimensional mapping technologies, including scanning hyperspectral imaging, due to the inherent complexity and heterogeneity of biological samples. Scientists from the Berkeley Synchrotron Infrared Structural Biology Imaging Program (BSISB) reviewed the history of adaptive sampling algorithms and surrogate modeling and summarized recent implementations of autonomous adaptive data acquisition (AADA) methods to benefit scanning hyperspectral imaging. IR hyperspectral imaging of biological systems is a noninvasive and label-free method for studying the time-resolved spatiochemical evolution of living biological systems. Using traditional approaches, staff scientists supporting user research activities at synchrotron facilities devote significant time to identifying regions of interest in user samples, which limits beamline efficiency and the number of users able to access the facilities. With advances in AADA, the time required to complete a user’s full data collection protocol can be significantly reduced by employing the system’s ability to autonomously and adaptively locate and analyze sample features of interest. The approach enhances overall efficiency and increases the number of users served. Advances in autonomous experimentation—closely linked to the original concept of artificial intelligence—have traditionally been limited by two recurring primary challenges. The first is shifts in distribution due to discrepancies between training datasets and conditions during testing or deployment. The second is difficulties in transfer learning in which information gleaned from a previously learned task must be transferred to a new task. The simplest solution is to separate the autonomous experimentation workflow into distinct, actionable components individually optimized for a user’s specific structural and functional goals. BSISB scientists conceptualize AADA as a flexible, tunable, “smart” building block framework that can be embedded into scanning hyperspectral imaging workflows to obtain high resolution and high precision chemical information from samples. | |||
| 01/18/2023 | Unelectrified Aerosol-Producing Device for Biological Studies | Berkeley Synchrotron Infrared Structural Biology Imaging Program | Cryo-Electron Microscopy, Solution X-ray Scattering, Synchrotron Infrared Hyperspectral Imaging, X-ray Absorption and Emission Spectroscopy, X-ray Fluorescence Imaging, X-ray Macromolecular Crystallography | Advanced Light Source | Molecular Structure, Cell and Tissue Structure, Chemical and Elemental Information |  The “whipping jet” device delivers a highly reproducible two-dimensional array of droplets of various sizes using a fine-scale microfluidic nozzle. [Reprinted from Narayanasamy et al., Copyright 2023, with permission from Elsevier.] The novel device has the potential to impact many fields of basic and applied research including structural biology, carbon capture, and climate science. Microdroplet production can also benefit sample environment instrumentation such as mass spectrometry, X-ray free-electron lasers (XFELs), synchrotrons, and cryo-electron microscopy (cryo-EM) used in, for example, biomacromolecule characterization. | |||
| 02/01/2023 | Bacterial Mercury Methylation Enhanced by Mixed Thiols | Structural Molecular Biology Resource | X-ray Absorption and Emission Spectroscopy | Stanford Synchrotron Radiation Lightsource | Chemical and Elemental Information | Results indicate that the effects of thiols on Hg methylation are more complex than previously known. Mercury complexed individually with the thiols cysteine, dithiol 2,3-dimercaptopropanesulfonate (DMPS), or 2,3-dimercaptosuccinic acid (DMSA) at low concentrations found in the environment strongly inhibited Hg methylation. Yet a mixture of cysteine with either DMSA or DMPS increased methylation rates by 1.5x-3.5x over the no-thiol control. Spectroscopic analyses indicated the formation of mixed Hg-thiolate complexes with coordination numbers of 3 or 4, which likely facilitated exchange of Hg with cells and its uptake and internal transfer to the proteins that carry out methylation. This foundational study underscores the need for detailed spectroscopic studies of environmental pollutants and biological controls that could act as natural pollutant remediation processes. | Inorganic mercury (Hg) released into the environment by humans is converted to the toxin monomethylmercury (MgHg) by certain anaerobic microorganisms. MeHg is present in waterways and bioaccumulates through aquatic food chains, posing a risk to wildlife and human health. Thus, it is important to understand biological control of environmental Hg transformation. Researchers at Oak Ridge National Laboratory (ORNL) and the Stanford Synchrotron Radiation Lightsource (SSRL) investigated how the presence of different low-molecular-weight thiol compounds, known to either inhibit or enhance Hg methylation, influence rates and extent of Hg methylation by the model methylating bacterium Geobacter sulfurreducens PCA (PCA). | Extended X-ray absorption fine structure (EXAFS) and X-ray absorption near edge structure (XANES) spectroscopy were performed at SSRL beamline 7-3 on mixtures of low-molecular-weight thiols complexed with Hg to determine potential mechanisms of complexation. Spectroscopy data were complemented by density functional theory calculations also conducted at SSRL, as well as ultraviolet-visible (UV-vis) and Raman spectroscopies at ORNL. | |
| 09/19/2022 | Viral Gene Products May Contribute to Soil Carbon Cycling | Structural Molecular Biology Resource | X-ray Macromolecular Crystallography | Environmental Molecular Sciences Laboratory, Stanford Synchrotron Radiation Lightsource | Molecular Structure | Metagenomics is unearthing the previously hidden world of soil viruses. Many soil viral sequences in metagenomes contain putative auxiliary metabolic genes (AMGs) unassociated with viral replication. Using X-ray macromolecular crystallography to visualize the atomic structure of proteins expressed by several viral AMGs, scientists established that the proteins are in fact functional and active. The researchers expressed and functionally screened AMGs that potentially encode chitosanase enzymes that metabolize chitin—an abundant soil carbon polymer found in insect exoskeletons and fungal cell walls. One expressed protein showing endo-chitosanase activity, called viral chitosanase (V-Csn), was crystalized and structurally characterized at ultra-high resolution. Specifically, researchers irradiated fragile crystallized protein samples with high-brightness X-rays generated by the Stanford Synchrotron Radiation Lightsource (SSRL) beamline 12-2. The V-Csn structure provides details about its active site and, together with structure models determined using the protein structure prediction software AlphaFold, facilitates understanding of substrate specificity and enzyme mechanism. The findings support the hypothesis that soil viruses contribute auxiliary functions to their hosts and may play a role in critical soil processes like chitin decomposition and carbon cycling. | |||
| 11/17/2022 | Direct Visualization of Fungal Mineral Transport | Structural Molecular Biology Resource | X-ray Fluorescence Imaging | Environmental Molecular Sciences Laboratory, Stanford Synchrotron Radiation Lightsource | Chemical and Elemental Information | Fungal species are foundational members of soil microbiomes, where their contributions in accessing and transporting vital nutrients is key to community resilience. The newly developed mineral-doped soil micromodel platform directly probes fungal growth using spatially resolved optical and chemical imaging methodologies. Results demonstrate that mineral presence is required for fungal thigmotropism—the turning or bending of an organism in response to contact with an object—around obstacles and through soil-like pore spaces. This is related to fungal transport of potassium and corresponding speciation from potassium-rich minerals. The findings provide visual mechanistic evidence of hyphal transport of mineral-derived nutrients under nutrient and water stresses. | Soil fungi facilitate translocation of inorganic nutrients from soil minerals to other microorganisms and plants. This ability is particularly advantageous in impoverished soils because fungal mycelial networks can bridge otherwise spatially disconnected and inaccessible nutrient hotspots. However, the molecular mechanisms underlying fungal mineral weathering and transport through soil remains poorly understood. This is primarily due to the lack of a platform for spatially resolved analysis of biotic-driven mineral weathering. Scientists addressed this knowledge gap by developing a mineral-doped soil micromodel platform to study mineral weathering mechanisms. The platform demonstrates fungal bridging of carbon hotspots and hyphal transport of mineral-derived nutrients. | Researchers used X-ray fluorescence (XRF) imaging and X-ray absorption near edge structure (XANES) spectroscopy at the Stanford Synchrotron Radiation Lightsource (SSRL) beamline 14-3 to measure potassium uptake in fungal hyphae. Specifically, they measured how organic acids exuded from Fusarium sp. Strain DS 682, a representative common saprotrophic soil fungus, chelate and uptake potassium from mineral interfaces. The measurements were complemented using mass spectrometry imaging techniques at the Environmental Molecular Sciences Laboratory (EMSL). | |
| 04/01/2022 | Mechanisms of Bacterial Iron Metabolism | Structurally Integrated Biology for the Life Sciences | Solution X-ray Scattering | Advanced Light Source, Advanced Photon Source | Molecular Structure | Iron (Fe) is an essential element for nearly all organisms, and under anoxic and/or reducing conditions, Fe2+ is the dominant form of iron available to bacteria. The ferrous iron transport (Feo) system is the primary Fe2+ import machinery in prokaryotic organisms, and two constituent proteins (FeoA and FeoB) are conserved across most bacterial species. However, how FeoA and FeoB function relative to one another remains enigmatic. Using a combination of structural modeling, SAXS, and hydrogen-deuterium exchange mass spectrometry, researchers showed that FeoA and FeoB interact in a nucleotide-dependent manner and mapped the protein-protein interaction interface. The results provide insight into mechanisms of bacterial iron import and utilization. The study also helps define a macromolecular switch that could be tuned to control iron utilization by microbes. As iron is a key constituent of many redox-sensitive reactions within microbes, the mechanism is fundamental to many systems within DOE mission space. | |||
| 04/21/2022 | Protein Design for Genetically Encodable Nanomachines | Structurally Integrated Biology for the Life Sciences | Cryo-Electron Microscopy, Solution X-ray Scattering | Advanced Light Source | Molecular Structure | Natural molecular machines contain protein components that undergo motion relative to each other. Designing such mechanically constrained nanoscale protein architectures is an outstanding challenge for computational protein design. Scientists explored the de novo construction of protein machinery from designed axle and rotor components that display internal cyclic or dihedral symmetry. Small-angle X-ray scattering (SAXS), at the DOE-supported Structurally Integrated Biology for the Life Sciences resource (SIBYLS), was used to screen nearly 100 axle and rotor constructs that would eventually be assembled to form axle-rotor systems (see figure). Once assembled, cryo-EM was used to verify that the constructs matched the design models. This combined approach confirmed that the axle-rotor systems assembled in vitro and in vivo as designed. Achieving construction of mechanical systems with internal degrees of freedom is a step toward the design of genetically encodable nanomachines. These proof-of-concept axle-rotor assemblies demonstrate that protein nanostructures with internal mechanical constraints can now be systematically designed. Key to this advance is the ability to computationally design protein components with complex complementary shapes, symmetries, and topologies, such as high-aspect-ratio dihedral axle structures (D2 homotetramers to D8 homo-16-mers) with oligomerization states and sizes considerably larger than previously designed. Efficient techniques like SAXS are necessary to validate computational design. Studying the assembly of shape-complementary homo-oligomeric components into higher-order hetero-oligomeric structures with internal degrees of freedom provides insight into further design of complex protein nanomachines. | |||
| 10/14/2022 | Modeling the Partitioning of Amphiphilic Molecules and Cosolvents in Biomembranes | Center for Structural Molecular Biology | Small-Angle Neutron Scattering | Spallation Neutron Source/High Flux Isotope Reactor | Molecular Structure | A primary limitation to fermentation processes is cosolvent and end-product cytotoxicity. Industrial solvents used in lignocellulosic pretreatment and aliphatic fermentation end-products interact with various cellular systems, impacting cell growth and yield in biofuel-producing microbes. The partitioning of small amphiphilic molecules into lipid bilayers from aqueous solution and the corresponding effect on bilayer structure is therefore of fundamental biophysical interest and of significant practical importance. Small-angle neutron scattering (SANS) is a key method for studying lipid and polymer bilayer structures. In this study, researchers developed a model for analyzing SANS measurements of solvent partitioning in lipid membranes. This is important for quantifying changes in the lipid bilayer structure of cells when exposed to biofuels and other fermentation products such as ethanol, butanol, or acetic acid. Existing models for interpreting SANS data of lipid membranes have struggled to reliably determine the molecular details of cosolvent partitioning into different regions of the lipid bilayer and changes to the bilayer structure. To address this, researchers developed a model of a bilayer structure with a two-term partition constant accounting for the localization of the cosolvent within the bilayer. The new model divides the lipid bilayer into three slabs: two for the head groups on either end of the bilayer and a single slab for the central acyl tail section. This model was successfully applied to SANS measurements of lipid vesicles in the presence of tetrahydrofuran (THF), yielding structural information for the bilayer and information about THF partitioning into the bilayer. Model assumptions were evaluated by molecular dynamics simulations. The approach yielded estimates of the partition coefficient for THF in 1,2-dimyristoyl-sn-glycero-3-phosphocholine at 35°C, along with an estimate of the fraction of THF residing in the hydrophobic core of the membrane. This analysis approach can be applied to many other bilayer/amphiphile interactions. The code needed to implement the model as an algorithm for scattering data has been added to the SASView software suite for small angle scattering data analysis. | |||
| 10/01/2022 | Detergent-Free Method for Photosystem I Stabilization in Structural Studies | Center for Structural Molecular Biology | Small-Angle Neutron Scattering, Solution X-ray Scattering | Spallation Neutron Source/High Flux Isotope Reactor | Molecular Structure | Oxygenic photosynthesis is the process by which plants, algae, and cyanobacteria convert sunlight into chemical energy, fueling all life on earth. It is performed by four membrane-associated protein complexes working in concert: photosystem II (PSII), cytochrome b6f, photosystem I (PSI), and ATP synthase. The oligomeric states of the individual domains of these protein complexes are relatively similar across kingdoms, except for PSI. PSI exists as a trimer in cyanobacteria and as a monomer in plants and algae. More recently, a tetrameric form has been found in heterocyst-forming cyanobacteria. This dynamic evolutionary history has spurred questions about how structural differences affect the overall function or activity of PSI in vivo. Further, detergent-based methods used to extract PSI from its thylakoid membrane environment for in vitro studies may alter its activity by modifying PSI interactions with thylakoid lipids that determine its function and structural integrity. Researchers tested an alternative to detergent solubilization to isolate trimeric PSI from the cyanobacterium Thermosynechococcus elongatus in which styrene maleic acid (SMA) copolymers were used to produce membrane protein-containing nanodics—model membranes for solubilizing and studying membrane proteins. The nanodics, called SMA lipid particles (SMALPs), retained the native lipid environment and preserved native protein function. Small-angle X-ray scattering (SAXS) and neutron scattering (SANS) were used to determine the size and shape of the particles in their fully solvated state. The particle’s proteolipid core and detergent shell or copolymer belt were interrogated separately using contrast variation, a capability unique to SANS. At ~1.5 MDa, PSI-SMALP is the largest SMALP yet isolated. SANS with contrast variation showed, for the first time, a ~1 nm SMA copolymer belt surrounding PSI, explaining the increased energy transfer observed in PSI embedded in SMALPs compared to detergent micelles. SMA copolymers hold great promise for stabilization of membrane proteins for structural studies. This study provides a structural basis for the observed increase in energy transfer and charge separation in PSI-SMALPs compared to detergent-isolated PSI complexes, highlighting the importance of a native lipid environment for maintaining PSI activity in vitro. | |||
| 07/05/2022 | Characteristics of Switchgrass Lignin Extracted by Different Solvent Pretreatments | Center for BioMolecular Structure | Solution X-ray Scattering | National Synchrotron Light Source II | Molecular Structure | Biomass recalcitrance due to the complex hierarchical structure of plant cell walls presents a challenge in biorefinery systems which seek to maximize biomass conversion to high-value products. Two approaches in biomass conversion that have succeeded in modifying the structure of lignocellulose to improve enzymatic deconstruction include thermochemical pretreatment and genetic modification. However, the characteristics of solubilized lignin resulting from different genotypes pretreated with different methods have not been extensively investigated. Organosolv pretreatments, which employ an organic solvent to solubilize the lignin and hemicellulose components of biomass, have demonstrated the ability to both maximize sugar release for downstream processing to fuels and also to enable extraction of relatively high purity lignin suitable for making bioproducts. In this study, researchers compared three organosolv pretreatment systems—ethanol, tetrahydrofuran (THF), and γ-valerolactone (GVL)—on three switchgrass genotypes: wildtype (WT) and two transgenic switchgrasses genetically modified via down-regulation of the caffeic acid/5-hydroxyconiferyl aldehyde O-methyltransferase gene (referred to as COMT) and overexpression of the MYB4 gene (referred to as MYB). A fundamental understanding of the impacts of organic solvents and reaction conditions on the structure and properties of switchgrass lignin helps develop and optimize lignin-targeting organosolv pretreatments. All organosolv pretreatments reduced the molecular mass of lignins, but to different degrees. THF-pretreated transgenic lignin retained the highest molecular mass, more β-O-4 linkages, and a higher aliphatic hydroxyl content than ethanol and GVL pretreated lignins. Ethanol pretreatment produced the greatest decrease in mass (∼90%) and near-complete removal of β-O-4 linkages. The abundant free phenolic hydroxide groups found in ethanol-pretreated lignin may make it suitable for development of antioxidant products, whereas GVL and THF lignins which retained more than half of their β-O-4 linkages may be more suitable for depolymerization to mono-aromatic compounds. Detailed structural information about the extracted lignins was also provided by synchrotron small-angle X-ray scattering (SAXS) using the LiX beamline at the National Synchrotron Light Source II (NSLSII). The nanostructure observations revealed that ethanol pretreatment produced the smallest lignin particles from WT switchgrass. These analyses complemented insights from nuclear magnetic resonance and molecular dynamics simulations. A correlation was found between the molecular mass reduction of lignin molecules in the experiments and the number of hydrogen bonds between lignin and the organic solvents as calculated in the MD simulation, suggesting a connection between the depolymerization of lignin and its ability to form a hydrogen bond with the organic solvents. | |||
| 06/11/2021 | Evolving a Virus-Like Capsid from a Bacterial Protein | Center for BioMolecular Structure | X-ray Footprinting | National Synchrotron Light Source II | Molecular Structure | Using directed evolution, an international team of scientists have converted a bacterial enzyme into an artificial protein shell that encapsulates the enzyme’s own encoding RNA. Such capsids have previously only been seen in viruses, which use them as part of their replication machinery. Understanding how viral capsids evolved as a system for infecting cells with viral genetic material may help researchers design novel delivery mechanisms for gene therapies. The team of scientists investigated the evolutionary history of how viruses build a protein capsid to protect their genetic material. Then they engineered an artificial capsid evolution system that started with a nucleocapsid derived from Aquifex aeolicus lumazine synthase, a bacterial enzyme that naturally forms 60-subunit nanocontainers. During artificial evolution, the nucleocapsid underwent a series of structural modifications, becoming much larger and more efficient at RNA uptake. Among other techniques, the researchers used X-ray footprinting at the X-ray Footprinting of Biological Materials (XFP) beamline at the National Synchrotron Light Source II (NSLS-II), which enabled a view inside the evolved capsids and insight into how they encapsidate RNA. The XFP beamline operates in close association with the BER-supported Center for BioMolecular Structure. | |||
| 10/08/2022 | XMIDAS Software Enables Chemical Analysis from Nanoscale Spectroscopy Data | Center for BioMolecular Structure | X-ray Absorption and Emission Spectroscopy, X-ray Fluorescence Imaging | National Synchrotron Light Source II | Chemical and Elemental Information | Synchrotron-based X-ray spectromicroscopy tools are widely used to understand the chemistry and morphology of complex material systems owing to their high penetration depth and sensitivity. High-resolution scanning X-ray spectromicroscopy expands the spectroscopy toolbox into the nanoscopic scale, but additional tools are needed to process the resulting multi-dimensional spectromicroscopy data. X-ray Multimodal Image Data Analysis Software (XMIDAS) is an open-source python package for analysis of spectromicroscopy data from both image and spectrum representations for nanoscale and microscale chemical imaging. XMIDAS is publicly distributed with the help of scientists at the Data Science & Systems Integration Program at Brookhaven National Laboratory’s (BNL) National Synchrotron Light Source II (NSLS-II). A key motivation behind the software development project, conducted in collaboration with BNL’s Center for BioMolecular Structure and NSLS-II’s Imaging and Microscopy Program, was to create a user-friendly tool to extract meaningful chemical information from high-dimensional (4D+) spectromicroscopy data. The program combines conventional data processing workflows with well-established machine-learning tools in an easy-to-use graphical user interface environment. Researchers used a combination of nanoprobe-based X-ray fluorescence spectromicroscopy (nano-XRF) X-ray absorption near edge structure spectromicroscopy (nano-XANES), and differential phase-contrast imaging to probe elemental and chemical state information of aggregate samples, and then used XMIDAS to visualize the complete chemistry of localized nanostructures. The optimized data-reduction strategies and tool development facilitate the analysis of complex biological and environmental samples at both micro- and nanoscales using X-ray spectromicroscopy techniques. | |||
| 08/26/2022 | RuBisCO Evolution Enabled by Structural Plasticity | Structurally Integrated Biology for the Life Sciences | Solution X-ray Scattering, X-ray Macromolecular Crystallography | Advanced Light Source | Molecular Structure | Researchers from Lawrence Berkeley National Laboratory studying the evolution of Ribulose-1,5-bisphosphate carboxylase-oxygenase (RuBisCO) reveal that the enzyme’s structural plasticity underlies its rich diversity of molecular assemblies. RuBisCO, the world’s most abundant protein, catalyzes the first step in carbon fixation. To better understand the phylogenetic distribution of the oligomeric states of form II RuBisCO, found in bacteria and certain photosynthetic microbes, the researchers structurally characterized 28 candidates spread across the phylogeny using traditional macromolecular X-ray crystallography (MX) and small-angle X-ray scattering (SAXS). SAXS offers lower resolution than MX but can take snapshots of proteins in their native form when suspended in solution. Combining this structural data with respective protein-coding gene sequences enabled the team to also perform ancestral sequence reconstruction. The reconstruction suggested that form II RuBisCO structure has remained plastic throughout its evolutionary history. To further test the theory that structural plasticity can give rise to new forms, the researchers performed mutational experiments, finding that functional changes in oligomerization could be produced with surprisingly few mutations. In some cases, the new RuBisCO assemblies exhibited improved affinity for their target molecule, carbon dioxide. The results suggest that form I RuBisCO engineering efforts in plants likewise may not need to focus exclusively on the enzyme’s active site to develop more productive and resource-efficient crops. X-ray scattering experiments were performed using the SIBYLS beamline at the Advanced Light Source and X-ray crystallography was performed at the Berkeley Center for Structural Biology. Research was also supported by the DOE Joint BioEnergy Institute and the Integrated Diffraction Analysis Technologies (IDAT) program. | |||
| 04/07/2022 | Soft X-ray Tomography Captures Mesoscale Organelle Interactions | National Center for X-Ray Tomography | Soft X-ray Tomography | Advanced Light Source | Cell and Tissue Structure | Inter-organelle interactions are a vital part of normal cellular function but are difficult to quantify due to the range of scales encountered in cell biology and the limitations of traditional imaging approaches. Soft X-ray tomography (SXT) can rapidly map ultrastructural reorganization and inter-organelle interactions in intact cells by taking advantage of the naturally occurring, differential X-ray absorption of carbon-rich compounds in each organelle. As an example, SXT was used to map the spatiotemporal evolution of insulin vesicles and their co-localization and interaction with mitochondria in pancreatic beta cells during insulin secretion and in response to different stimuli. The technique enabled quantification of changes in the morphology, biochemical composition, and relative position of mitochondria and insulin vesicles. The findings highlight the importance of comprehensive and unbiased mapping at the mesoscale to characterize cell reorganization that would be difficult to detect and quantify with other existing methodologies. The same technique can be useful for quantifying morphological changes inside a microbial cell in response to genomic manipulations or other perturbations. | |||
| 01/11/2022 | Biocatalyst Inspiration: Specificity and Selectivity of Rieske Oxygenases | Structural Molecular Biology Resource | X-ray Macromolecular Crystallography | Stanford Synchrotron Radiation Lightsource | Molecular Structure | Rieske non-heme iron oxygenases represent one of nature’s solutions for performing precise site-selective C–H bond functionalization reactions by exploiting the reactivity of iron. Thus far, only a handful of Rieske oxygenases have been structurally characterized and remarkably little information exists regarding how these enzymes use a common architecture and set of metallocenters to facilitate a diverse range of reactions. To investigate the architectural parameters that dictate the substrate specificity and site-selectivity of a Rieske oxygenase catalyzed reaction, scientists focused on two Rieske oxygenases, SxtT and GxtA, which are involved in the biosynthesis of paralytic shellfish toxins. They used macromolecular X-ray crystallography to detail how SxtT and GxtA use different protein regions to influence the site-selectivity of their catalyzed monohydroxylation reactions and the location of a substrate access tunnel to the active site. The structural information allowed for the identification of six residues distributed between three regions of SxtT that together control the selectivity of the C–H hydroxylation event. Substitution of the residues produced a SxtT variant fully adapted to exhibit the non-native site-selectivity and substrate scope of GxtA. Importantly, the selectivity regions are conserved in other structurally characterized Rieske oxygenases, providing a framework for predictively repurposing and manipulating Rieske oxygenases as biocatalysts. | |||
| 03/25/2021 | Understanding How Bacteria Degrade N-Glycan | Structural Molecular Biology Resource | X-ray Macromolecular Crystallography | Stanford Synchrotron Radiation Lightsource | Molecular Structure | N-glycosylation is a fundamental protein modification found in both eukaryotes and archaea, but rarely bacteria. Despite lacking N-glycans, many commensal and pathogenic bacteria have developed mechanisms to degrade scavenged N-glycans for a variety of functions including nutrient acquisition; they express enzymes that cleave the glycan chain’s core. If the glycan is attached to a protein, it is first removed before cleavage. Although much is known about many enzymes responsible for N-glycan degradation, the enzymes involved in cleaving the N-glycan core have only recently been discovered. Thus, some of the structural details have yet to be characterized and little is known about their full distribution among bacterial strains. Researchers used macromolecular X-ray crystallography to elucidate the structure of the active site of a family of glycoside hydrolases—enzymes responsible for glycan core degradation—from the soil bacterium Streptomyces cattleya and the gut bacterium Bifidobacterium longum. The study provides understanding of how these bacteria degrade glycan, pointing to an efficient mechanism for soil microbes to recycle decomposing biomass. | |||
| 11/04/2021 | Potentially Customizable Molecular Assembly Line | Stanford-SLAC Cryo-EM Center, Structural Molecular Biology Resource | Cryo-Electron Microscopy, X-ray Macromolecular Crystallography | Advanced Light Source, Advanced Photon Source, Stanford Synchrotron Radiation Lightsource | Molecular Structure | Researchers used X-ray macromolecular crystallography (MX) and cryogenic electron microscopy (cryo-EM) to describe the atomic structure of a modular enzyme—a polyketide synthase—intact and at different stages of its reaction cycle. The experiments revealed that the enzyme, a molecular assembly line producing common antibiotics, is composed of two interacting yet asynchronously-operating reaction chambers. A high-resolution structure of an intact polyketide synthase module had remained elusive for over 30 years since the enzyme’s discovery. The team used multiple beamlines at the Stanford Synchrotron Radiation Lightsource, the Advanced Light Source, and the Advanced Photon Source over a period of three years to elucidate its structure and mechanism. The unprecedented detail will inform future studies aimed at engineering polyketide synthases to produce “green” antibiotics, pharmaceuticals, and novel biofuels. | |||
| 04/05/2022 | How a Soil Microbe Could Rev Up Artificial Photosynthesis | Structural Molecular Biology Resource | X-ray Macromolecular Crystallography | Advanced Photon Source, Stanford Synchrotron Radiation Lightsource | Molecular Structure | A team of researchers from DOE’s SLAC National Accelerator Laboratory and Joint Genome Institute, the Max Planck Institute for Terrestrial Microbiology in Germany, and the University of Concepción in Chile has discovered how a bacterial enzyme carries out a key step in carbon fixation 20 times faster than plant enzymes during photosynthesis. The findings provide a basis for engineering highly efficient CO2-fixing enzymes for bioenergy and bioproduct applications. The team used a combination of ambient-temperature X-ray free electron laser (XFEL) and cryogenic synchrotron experiments to study the structural organization of enoyl-CoA carboxylase/reductase (ECR) from Kitasatospora setae. The approach revealed that rather than grabbing carbon dioxide molecules and attaching them to biomolecules one at a time, the ECR from K. setae consists of pairs of molecules that work in sync to get the job done faster. One member of each enzyme pair opens wide to catch a set of reaction ingredients while the other closes over its captured ingredients and carries out the carbon-fixing reaction; then, they switch roles in a continual cycle. A single spot of molecular “glue” holds each pair together so they can alternate opening and closing in a coordinated way, while a twisting motion helps shuttle ingredients and finished products in and out of the pockets where the reactions take place. When both “glue” and “twist” are present, the carbon-fixing reaction proceeds 100 times faster than without them. | |||
| 12/16/2021 | Aggregates Formed During Biomass Pretreatment Stem from Both Hemicellulose and Lignin | Center for Structural Molecular Biology | Small-Angle Neutron Scattering | Spallation Neutron Source/High Flux Isotope Reactor | Molecular Structure | Production of second-generation bioethanol from lignocellulosic biomass requires pretreatment to open the plant cell wall structure and improve enzyme access. But lignin aggregates formed during dilute acid pretreatment (DAP) of biomass are known to contribute to lower sugar yields for biofuel production. To understand the role of noncellulosic switchgrass polymers on the overall efficiency of pretreatment, researchers used time-resolved in situ small-angle neutron scattering (SANS) to investigate real-time structural changes in the noncellulosic polymers of switchgrass plant cell walls during DAP. They found that hemicellulose forms aggregates between 80-130°C, whereas at higher temperatures lignin-derived aggregates are observed. The study provides the first direct evidence that early stages of pseudo-lignin aggregate formation within plant cell walls originate from hemicellulose, not just lignin, during thermochemical pretreatment and that both contribute to decreased enzyme accessibility and biomass recalcitrance.
Trends in the growth of lignin (blue) and pseudo-lignin (red) aggregate particles during dilute acid pretreatment. [Reprinted with permission from Yang, et al. 2022.] | |||
| 09/14/2021 | A Tunable Plant Pseudoenzyme-Enzyme Complex Regulates B6 Production | Cryogenic Transmission Electron Microscopy at EMSL | Cryo-Electron Microscopy | Environmental Molecular Sciences Laboratory | Molecular Structure | Pseudoenzymes—catalytically deficient or inactive homologs of active enzymes—can act as regulatory elements, often by pairing with their active counterparts. However, the high structural similarities between pseudoenzymes and canonical enzymes create challenges for unraveling their mechanisms of action using classical molecular and biochemical characterization approaches. Vitamin B6 biosynthesis in plants is one example where pseudoenzyme-enzyme pairing, called heterocomplexation, between the enzyme PDX1.3 and the pseudoenzyme PDX1.2 impact overall activity, yet the mechanism remained unclear. Researchers from four institutions, including the Environmental Molecular Sciences Laboratory (EMSL), used a unique combination of cryo-EM, cell-free expression, activity assays, and native mass spectrometry to determine the atomic structure and manipulate the degree of pseudoenzyme-enzyme pairing. The various pseudoenzyme-enzyme pair associations exhibited variable stoichiometry that was tunable, solving an open question on the mechanism of heterocomplex assembly and activity regulation. Instead of forming discrete rings of pure pseudoenzyme or enzyme hexamers that can interact in a ring-ring association, the results revealed that the pseudoenzyme and enzyme subunits interact both within and between each ring layer. | |||
| 02/22/2021 | Repurposing Cancer and Seizure Medications to Aid Fight Against COVID-19 | Structural Biology Center | X-ray Macromolecular Crystallography | Advanced Photon Source | Molecular Structure | Two teams of researchers used resources at the Structural Biology Center at Argonne National Laboratory’s (ANL) Advanced Photon Source to investigate two drugs that may be repurposed to fight SARS-CoV-2, the virus that causes COVID-19. In research led by Andrzej Joachimiak of ANL and the University of Chicago, along with ANL protein crystallographer Youngchang Kim and University of Chicago structural biologist Natalia Maltseva and colleagues, the team found that tipiracil, a drug used to treat colorectal cancer, can inhibit the action of one of the main proteins that comprise SARS-CoV-2. They used high-powered X-ray beams to study one of the virus’s proteins, called Nsp15. In a separate study, scientists from Yale University used the Structural Biology Center resource to study the structure of perampanel, an anti-seizure medication, as a starting point for SARS-CoV-2 inhibitor design. Modifying perampanel to create new configurations of the drug gave researchers new molecules that were effective against the virus. The new molecules would be used along with remdesivir, a current therapeutic agent for COVID-19. | |||
| 06/11/2020 | Structural Basis for Neutralization of Betacoronaviruses by Llama Antibodies | Structural Biology Center | X-ray Macromolecular Crystallography | Advanced Photon Source | Molecular Structure | Coronaviruses use a large envelope protein called spike (S) to engage host cell receptors and catalyze membrane fusion. S proteins therefore represent a vulnerable target for therapeutics. Scientists isolated single-domain antibodies (VHHs) from a llama immunized with prefusion-stabilized coronavirus spike proteins. The VHHs neutralized MERS-CoV and SARS-CoV-1 S pseudotyped viruses. Crystal structures of the VHHs bound to their respective viral targets revealed two distinct epitopes, but both VHHs interfered with receptor binding. There was also cross-reactivity between the SARS-CoV-1 S-directed VHH and SARS-CoV-2 S; the cross-reactive VHH neutralized SARS-CoV-2 S pseudotyped viruses. The data provide a molecular basis for the neutralization of pathogenic betacoronaviruses by VHHs and suggest that the molecules may serve as useful therapeutics during coronavirus outbreaks. | |||
| 08/19/2021 | Existing Drug Inhibits SARS-CoV-2 Replication | Structural Biology Center | X-ray Macromolecular Crystallography | Advanced Photon Source | Molecular Structure | Inside host cells, the RNA genome of severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) is translated into two polyproteins that are cleaved to give the individual viral proteins. The main viral protease, known as Mpro or 3CLpro, plays a key role in these cleavages, making it an important drug target. Drayman et al. identified eight drugs that target 3CLpro from a library of 1,900 clinically safe drugs. Because of the challenge of working with SARS-CoV-2, they started by screening for drugs that inhibit the replication of a human coronavirus that causes the common cold. They then evaluated the top hits for inhibiting SARS-CoV-2 replication and for inhibiting 3CLpro. Masitinib, a broad antiviral, inhibited the main proteases of coronaviruses and picornaviruses and was effective in reducing SARS-CoV-2 replication in mice. | |||
| 08/03/2021 | Deconstructing the Infection Machinery of SARS-CoV-2 | Center for Structural Molecular Biology, Structural Biology Center, Structurally Integrated Biology for the Life Sciences | Small-Angle Neutron Scattering, Solution X-ray Scattering, X-ray Macromolecular Crystallography | Advanced Light Source, Advanced Photon Source, Spallation Neutron Source/High Flux Isotope Reactor | Molecular Structure | The replication transcription complex (RTC) from the virus SARS-CoV-2 is responsible for recognizing and processing RNA for two principal purposes: copying viral RNA for propagation into new virus and ribosomal transcription of viral proteins. Scientists conducted a systematic structural investigation of three components of the RTC—Nsp7, Nsp8, and Nsp12—and solved high-resolution crystal structures of the Nsp7/8 complex, providing insight into the interaction between the proteins. The investigation involved a broad approach using a range of structural biology techniques, including small-angle x-ray and neutron solution scattering (SAXS and SANS) on all components and multiangle light scattering-coupled SAXS to identify which combination of components forms transient or stable complexes. Results indicated that individual Nsp7, Nsp8, and Nsp12 structures vary based on whether other proteins in their complex are present. Combining the determinations of crystal structure, atomic coordinates reported elsewhere, SAXS, SANS, and other biophysical techniques, the multidisciplinary research team provided insight into RTC assembly, mechanism, and potential avenues for disruption of the complex and its functions. | |||
| 05/12/2021 | Crystallography Confirms Drug-Protein Interaction that May Prevent Certain Breast Cancers | Structural Biology Center | X-ray Macromolecular Crystallography | Advanced Photon Source | Molecular Structure | Scientists used X-ray macromolecular crystallography at the Advanced Photon Source (APS), a U.S. Department of Energy Office (DOE) of Science User Facility at the DOE’s Argonne National Laboratory, to determine the structures of estrogen receptors on breast cancer cells in mice and the drug lasofoxifene, confirming that lasofoxifene binds to the receptors. The APS uses ultrabright X-rays to illuminate the structures of proteins like estrogen receptors, often looking to see if drug compounds attach themselves to these proteins. Structures were determined at the Structural Biology Center (SBC) and at the South-East Regional Collaborative Access Team beamlines. About 75% of breast cancers are estrogen receptor positive, meaning that the hormone feeds tumor growth. Treatment with lasofoxifene outperformed fulvestrant, the current gold-standard drug, in reducing or preventing primary breast cancer tumor growth in the mice. The drug also was more effective at preventing metastasis to the lung, liver, bone, and brain — the four most common areas for breast cancer to spread. Human clinical trials are underway. | |||
| 04/17/2020 | Crystal Structures of SARS-CoV-2 Yield Understanding of Viral Replication Engine | Structural Biology Center | X-ray Macromolecular Crystallography | Advanced Photon Source | Molecular Structure | The rapid upsurge and proliferation of SARS-CoV-2 raised questions about how the virus became so much more transmissible than the SARS-CoV-1 and MERS-CoV coronaviruses. Although the viruses are similar, detailed information about SARS-CoV-2 protein structures and functions is needed to rapidly develop effective vaccines, antibodies, and antivirals. Scientists applied a high-throughput protein production and structure determination pipeline to produce SARS-CoV-2 proteins and structures. They report two high-resolution crystal structures of nonstructural protein 15 (Nsp15), an endoribonuclease conserved across coronaviruses that processes viral RNA to evade detection by host defense systems. The Nsp15 structure is very similar to the SARS-CoV-1 and MERS-CoV homologs, but shows some differences that may contribute to its greater virulence. | |||
| 05/01/2021 | Structural Plasticity of the Selectivity Filter in a Nonselective Ion Channel | Structural Biology Center | X-ray Macromolecular Crystallography | Advanced Photon Source | Molecular Structure | A crystallographic structure of a bacterial sodium-potassium ion channel (NaK) reveals conformational differences between the residues comprising the narrow pore within the channel, called the selectivity filter, and similar residues within selective potassium ion channels. NaK is a nonselective, monovalent cation channel that conducts sodium and potassium ions equally efficiently across the cellular membrane even though its selectivity filter differs by only one amino acid residue from filters that selectively conduct potassium ions. Potassium and sodium ions are highly abundant in biological systems, playing key roles in the electrical activity of cells. The crystallographic structure identifies a side-entry, ion conduction pathway for Na+ permeation that is unique to NaK. Nuclear magnetic resonance studies and molecular dynamics simulations confirmed the dynamical nature of the top part of the selectivity filter and the inner helix in NaK as also observed in the crystal structure. Taken together, the results indicate that the structural plasticity of the selectivity filter combined with the dynamics of the inner helix of NaK are vital for the efficient conduction of different ions through the non-selective NaK ion channel. | |||
| 10/05/2021 | Study Sheds Light on Photosynthesis in Iron-Low Leaves | Center for BioMolecular Structure | X-ray Fluorescence Imaging | National Synchrotron Light Source II | Chemical and Elemental Information | Researchers have identified how iron-deficient plants optimize photosynthesis to protect themselves from absorbing too much light, according to a study published in Proceedings of the National Academy of Sciences. Iron deficiency has many adverse effects on photosynthesis, which is not surprising given that 90% of foliar iron is found in the chloroplast. Although many iron-sparing mechanisms have been documented, a mechanistic understanding of how chloroplasts adapt to iron deficiency was lacking. The study demonstrated that the bHLH transcription factors ILR3 and PYE are required for photoprotection during iron deficiency. Changes in chloroplast morphology under the control of these transcription factors prevents the production of harmful reactive oxygen species and allows for repair of photosystem II. Understanding how plants adapt the photosynthetic machinery during iron deficiency may enable optimization of plant growth in soils where iron is not bioavailable. | |||
| 02/11/2021 | A Modular Approach to Designing Protein Assemblies | Structurally Integrated Biology for the Life Sciences | Solution X-ray Scattering | Advanced Light Source | Molecular Structure | Tailor-made nanostructures enable precise control over three-dimensional spatial arrangements and biochemical processes at the molecular level. Biological macromolecules, such as DNA and polypeptides, represent versatile, programmable biomaterials suitable for this purpose. In designing nanostructures, modularity is a commonly employed concept since it greatly simplifies the design process. In the protein realm, the modular paradigm is achievable by employing α-helical elements, such as coiled-coil (CC) units, as building modules. Scientists used a type of modular design based on pairwise-interacting CC units, called CC protein origami (CCPO), to create protein cages that self-assembled into oligomers. The conformation adopted by the protein cage in solution was examined using SAXS conducted at the SIBYLS beamline 12.3.1 at the Advanced Light Source. In addition, by introducing a protease cleavage site the researchers created a proteolysis-mediated conformational switch, demonstrating that polyhedral protein cages can be designed to transition between two structural states in response to an external cue. Self-assembly of CC-based nanostructures from several chains, similarly as in DNA nanotechnology, could facilitate the design of more complex protein assemblies and functionalities. | |||
| 01/19/2021 | Fusion Enzymes Improve Yield of Functionalized Terpenes | Structurally Integrated Biology for the Life Sciences | Solution X-ray Scattering | Advanced Light Source | Molecular Structure | Terpenes are a large class of natural products that can be used as precursors of fuel additives, fragrances, insecticides, pharmaceuticals, and other bio-based compounds. Enzymes such as cytochrome P450 can play an important role in the modification of terpenes essential for new bioactivities. However, the hydrophobicity and volatility of terpene molecules can limit the availability of the substrate around the enzyme and result in low enzymatic conversion during microbial production. In this study, researchers developed a strategy to improve the accessibility of terpene molecules for the P450 reaction by linking terpene synthase and P450 together into a synthetic fusion protein that draws P450 into closer contact with its terpene substrate. Several of the engineered fusion proteins they tested significantly increased the production of oxidized terpenes. Structural analysis of the fusion proteins was carried out using Size Exclusion Chromatography coupled to Small Angle X-ray Scattering (SEC-SAXS) at the SIBYLS beamline 12.3.1 at the Advanced Light Source. Analysis of positive and negative examples of the fusion strategy revealed key factors enabling structure-based prediction and evaluation of effective fusion enzymes. The results suggest that developing fusion enzymes of terpene synthase and P450 presents an efficient and widely applicable strategy for improving the biosynthetic titer of functionalized products from hydrophobic terpene intermediates. | |||
| 01/06/2021 | Designing a Controllable Self-Assembling Biomaterial | Structurally Integrated Biology for the Life Sciences | Solution X-ray Scattering | Advanced Light Source | Molecular Structure | The ability to manipulate the components of biomaterials and to control their assembly enables the design of materials with a wide variety of applications. Most previously described 2D protein materials, such as S-layers and de novo-designed arrays, primarily involve single protein components which spontaneously self-assemble, complicating characterization and repurposing for specific tasks. To address this problem, scientists have now developed a computational method to generate co-assembling binary layers by designing rigid interfaces between pairs of dihedral protein building blocks. And they used the method to design a hexagonal lattice. Investigation of the kinetics and assembly mechanism in vitro was conducted with high-throughput solution X-ray scattering (small-angle X-ray scattering; SAXS) at the SIBYLS beamline 12.3.1 at the Advanced Light Source. When combined at nanomolar concentrations, the proteins rapidly assembled into nearly crystalline micrometer-scale arrays almost identical to the computational design model in vitro and in cells without the need for a two-dimensional support. With this method, protein components can be readily functionalized and their symmetry reconfigured, enabling formation of ligand arrays with distinguishable surfaces. The resulting materials can impose order onto fundamentally disordered substrates such as cell membranes. | |||
| 02/03/2020 | Antibodies Elicited by Zika Vaccine Characterized | Center for BioMolecular Structure, Structural Biology Center | X-ray Macromolecular Crystallography | Advanced Photon Source, National Synchrotron Light Source II | Molecular Structure | A single dose of the experimental Zika virus vaccine, called ZPIV, given to a dengue-experienced individual boosted pre-existing flavivirus immunity and elicited protective cross-neutralizing antibody responses against both Zika and dengue viruses. The findings were based in part on research carried out at two U.S. DOE X-ray light sources: the Advanced Photon Source and the National Synchrotron Light Source II. After noting a significantly greater immune response in the dengue-experienced volunteer compared to dengue-naïve volunteers after vaccination, researchers isolated and characterized antibodies from the experienced volunteer. They found that one of the antibodies, MZ4, was particularly effective at protecting mice from infection by both Zika and dengue virus serotype-2 strain. In addition, vaccination of individuals in Puerto Rico with ZPIV who had prior flavivirus experience yielded similar cross-neutralizing potency after a single vaccination, highlighting the potential benefit of Zika vaccination in flavivirus-endemic areas. The study suggests that Zika virus vaccination can boost existing immune responses to dengue virus while generating potent Zika-neutralizing responses and may have unique potential as a preventative strategy in settings where both viruses are prevalent. | |||
| 09/24/2019 | Revealing a MERS-CoV Vulnerability | Structural Biology Center | X-ray Macromolecular Crystallography | Advanced Photon Source | Molecular Structure | Researchers used X-ray macromolecular crystallography to derive the molecular structure and functional characterization of G2, a neutralizing antibody targeting the spike glycoprotein of the Middle East respiratory syndrome coronavirus (MERS-CoV). MERS-CoV was first identified in June 2012 and has an estimated case fatality rate of 36%. The MERS-CoV spike protein comprises two subunits, S1 and S2. The S1 subunit mediates binding of the virus to the host cell receptor via its dipeptidyl peptidase-4 receptor (DPP4). Several neutralizing antibodies have been found that target either the N-terminal domain (NTD) or the receptor binding domain (RBD) of the S1 subunit, but those that target the former (S1-NTD) have not been well-characterized. Crystal structures of G2, alone and in complex with its S1-NTD target, were obtained at the U.S. DOE’s Advanced Photon Source along with biochemical, biophysical, and cell-based assays. The data reveal a site of vulnerability on S1-NTD and point to a neutralization mechanism whereby G2 inhibits the attachment of the MERS-CoV spike protein to the DPP4 receptor, preventing infection of host cells. The results increase the understanding of the viral attachment mechanism and may facilitate the development of S1-NTD-based vaccines against MERS-CoV. | |||
| 10/27/2020 | Potent Antibody Against SARS-CoV-2 | Structural Biology Center | X-ray Macromolecular Crystallography | Advanced Photon Source | Molecular Structure | Researchers employed high-brightness X-rays from the U.S. Department of Energy’s Advanced Photon Source (APS) to show how a monoclonal antibody from a COVID-19 survivor acts as a potent neutralizer of SARS-CoV-2. The antibody, CV30, binds to the receptor binding domain on the SARS-CoV-2 spike protein and induces it to shed its S1 subunit. Without the S1 subunit, which normally facilitates binding of the spike protein to the ACE2 receptor on human cells, the virus can no longer infect cells. CV30 also binds to a region of the ACE2 receptor that overlaps with the binding region of the spike protein, thereby blocking its access to the cell. CV30 may prove useful in the prevention or treatment of COVID-19. To find out, the antibody, along with other candidate proteins, needs to be tested pre-clinically and then in human trials. | |||
| 02/02/2021 | SARS-CoV-2 Enzyme Structure Reveals Potential Inhibitors | Structural Biology Center | X-ray Macromolecular Crystallography | Advanced Photon Source | Molecular Structure | Research at the U.S. Department of Energy’s Advanced Photon Source (APS) published in Nature Communications reports a high-resolution structure of Papain-like protease (PLpro), one of at least 29 different proteins encoded by SARS-CoV-2, the virus that causes COVID-19. PLpro is an enzyme responsible for releasing other proteins from a large protein complex and plays an important role in disrupting the host immune response. Structural, biochemical, and virus replication studies identified a set of chemical compounds capable of blocking PLpro activity in vitro. A subset of the compounds further demonstrated an ability to block SARS-CoV-2 from replicating in cell culture assays. Such inhibitors will need to undergo in vivo testing in animal models before being tested in human clinical trials, but the findings accelerate structure-based drug design efforts targeting PLpro to identify high-affinity inhibitors of clinical value. | |||
| 05/21/2021 | Essential Enzyme in Bacterial Protein Synthesis | Structural Biology Center | X-ray Macromolecular Crystallography | Advanced Photon Source | Molecular Structure | In recent work using the Structural Biology Center X-ray beamline 19-ID at the U.S. DOE’s Advanced Photon Source, researchers gained important insights into the structure and mechanism of phenylalanyl-tRNA synthetase, a critical enzyme involved in protein synthesis in Mycobacterium tuberculosis. Multi-drug resistant and extensively drug-resistant strains of M. tuberculosis have become a major global problem, increasing the need for new antibiotics. This has driven researchers to seek new bacterial targets for structure-based drug design. The work, published in Nucleic Acids Research, will form the basis of structure-based drug design efforts aimed at exploiting the unique features of the enzyme, a method that has been successful in developing pharmaceuticals against methicillin-resistant Staphylococcus aureus (MRSA) and some fungal infections. | |||
| 06/02/2021 | Complexation of Metals with Bacterial Methanobactins | Structural Molecular Biology Resource | X-ray Absorption and Emission Spectroscopy | Stanford Synchrotron Radiation Lightsource | Chemical and Elemental Information | The results lay the foundation for future work exploring the impact of methanobactins on the speciation and biogeochemical cycling of transition metals, including highly toxic methylmercury. | Spectroscopic techniques and time-dependent density functional theory (TD-DFT) calculations shed light on the structure and geometry of bacterial methanobactins binding to the transition metals Zn, Cd, and Hg. | Methanobactins are small peptides secreted by methanotrophic bacteria to facilitate their acquisition of Cu, an element they need to catalyze aerobic oxidation of methane to methanol. These methanobactins can also bind other trace metals, including the group 12 transition metals Zn, Cd, and Hg. Therefore, the structure of methanobactin-metal complexes has implications for the transport, bioavailability, and toxicity of trace metals in the environment. This study sought to understand the structure and geometry of methanobactins binding to Zn, Cd, and Hg. The complexation of these metals by methanobactin from Methylocystis sp. strain SB2 was studied using a combination of absorbance, fluorescence, extended x-ray absorption fine structure (EXAFS) spectroscopy, and TD-DFT calculations. Collectively, the results represent the first combined computational and experimental spectroscopy study of methanobactins and shed new light on molecular interactions and dynamics that characterize complexes of methanobactins with Group 12 transition metals. Research Details
| |
| 03/24/2021 | Deciphering the Mechanism of Enzymatic Methane Synthesis | Structural Molecular Biology Resource | X-ray Absorption and Emission Spectroscopy, X-ray Fluorescence Imaging | Stanford Synchrotron Radiation Lightsource | Chemical and Elemental Information | An unprecedented metalloenzyme structure containing nickel (Ni) is believed to initiate methanogenesis via a long-range electron transfer mechanism, expanding the understanding of metal-based catalysis in biology. | A solution to the substrate-binding conundrum in the mechanism of biological methane synthesis by methyl coenzyme M reductase (MCR). | Methane is the simplest organic compound with the highest energy content of any carbon-based fuel. Thus, understanding the biosynthesis of methane is imperative from basic energy, economic and environmental perspectives. This has led to extensive studies honing in on MCR and its role in catalyzing both the synthesis and anaerobic oxidation of methane. MCR is one of the few Ni-containing proteins in nature, and it is this Ni center that catalyzes the reaction of methyl-coenzyme M (CH3−SCoM) with coenzyme B (HSCoB) to form methane and the heterodisulfide CoMS−SCoB. Although significant mechanistic studies have been undertaken, none has successfully characterized binding of methyl-SCoM and CoMSSCoB with the active Ni(I) state. This study describes the coordination chemistry at the active Ni(I) site, elucidating a unique long-range electron transfer process in the MCR reaction mechanism. Clarification of Ni(I)–sulfonate binding requires the field to reassess the previously proposed mechanisms of this important reaction to consider long-range electron-transfer processes. Research Details
| |
| 10/30/2020 | Molecular Crystals Hybridized with Hydrogel Polymers Achieve Unprecedented Material Attributes | Structural Molecular Biology Resource | Solution X-ray Scattering, X-ray Macromolecular Crystallography | Stanford Synchrotron Radiation Lightsource | Molecular Structure | The ability to produce materials with desired mechanical and dynamic properties by deliberately designing their building blocks could enable a wide range of applications such as gas sorption and separation, sensing, and controlled release, among others. The transfer and amplification of atomic or molecular-level design to the macroscopic scale requires molecular building blocks to be organized and appropriately interconnected over multiple length scales. Crystalline materials provide a distinct advantage in this regard in that they are composed of only one or few components arranged periodically, possessing both short and long-range order to allow structural and mechanical coupling in a cooperative manner. However, the flexibility and adaptiveness of crystalline materials are rather limited, restricting the range of achievable macroscopic changes. Combining protein crystals, such as mesoporous ferritin, with hydrogel networks has resulted in hybrid materials called polymer-integrated crystals (PIX) which can undergo dramatic structural changes while maintaining crystalline periodicity and displaying efficient self-healing. But the uniform distribution and isotropic expansion/contraction of the lattice rendered the first-generation PIX materials isotropic, or lacking directionality. A research group from the University of California San Diego led by F.A. Tezcan showed that PIX materials can be patterned by the orientation and structural details of the distinct protein−protein interfaces in non-cubic ferritin lattices to display directional expansion and contraction and rapid bending motions, while retaining crystalline order. The process of crystal expansion and contraction was monitored in situ using small angle X-ray scattering measured at the Stanford Synchrotron Radiation Lightsource beamline 4-2, which confirmed that the cystalline lattice is intact throughout the process. Key to attaining anisotropic properties in PIX is the ability of the ferritin molecules to form lattices with distinct symmetries and protein−protein interfaces. These differences allowed the templation of alternatively-patterned hydrogel networks in situ, which ultimately enabled ferritin crystals that possess the same macroscopic morphologies to display orthogonally directed motions. Combined with the inherent chemical versatility and functions of proteins, such covalently hybridized PIX could offer a unique platform for the study of protein−polymer interactions and the development of biocatalytic and molecular encapsulation/delivery systems with tunable and responsive mechanical properties. For example, the hybridized PIX expand and contract anisotropically without losing crystallinity, undergo prompt bending motions in response to stimuli, and self-heal efficiently, capturing some of the essential features of sophisticated biological devices like skeletal muscles. | |||
| 01/07/2020 | Multistep Crystallization of Microbial S-Layers | Structurally Integrated Biology for the Life Sciences, Structural Molecular Biology Resource | Cryo-Electron Microscopy, Solution X-ray Scattering, X-ray Macromolecular Crystallography | Advanced Light Source, Stanford Synchrotron Radiation Lightsource | Molecular Structure | Many microbes assemble a crystalline protein layer on their outer surface as an additional barrier and communication platform between the cell and its environment. These surface layer (S-layer) proteins efficiently crystallize to continuously coat the cell, and this trait has been utilized to design functional macromolecular nanomaterials. A collaborative research team from Stanford University and the University of British Columbia, led by S. Wakatsuki, examined the structural basis for self-assembly of RsaA, a 98-kDa S-layer protein from Caulobacter crescentus, in vitro using a combination of time-resolved small angle x-ray scattering performed at SSRL’s beam line 4-2, circular dichroism spectroscopy, X-ray macromolecular crystallography, and a time course of cryogenic electron microscopy (Cryo-EM). The team established that the assembly pathway involves two domains serving distinct functions. The C-terminal crystallization domain forms the physiological 2-dimensional (2D) crystal lattice, but full-length protein crystallizes multiple orders of magnitude faster due to the N-terminal nucleation domain. Crystallization observations using a time course of cryo-EM imaging revealed a crystalline intermediate wherein N-terminal nucleation domains exhibit motional dynamics with respect to rigid lattice-forming crystallization domains. Dynamic flexibility between the two domains rationalizes efficient S-layer crystal nucleation on the curved cellular surface. The research demonstrates that a discrete nucleation domain is responsible for enhancing the rate of self-assembly, unveiling possible mechanisms to engineer kinetically controllable self-assembling 2D macromolecular nanomaterials. | |||
| 03/23/2020 | Antibiotics with a Built-in OFF Switch | Structural Molecular Biology Resource | Solution X-ray Scattering, Synchrotron Infrared Hyperspectral Imaging | Stanford Synchrotron Radiation Lightsource | Molecular Structure, Chemical and Elemental Information | Widespread use of antibiotics in healthcare and agriculture has led to their artificial accumulation in natural habitats. Emerging evidence illustrates a wide range of negative consequences of antibiotic waste accumulation in ecosystems. These include short-term and long-term adverse effects on the structure and function of microbial communities involved in biogeochemical cycling and organic matter degradation; contamination of water, plants, stockbreeding, and aquaculture products in the food chains; and the development of resistomes—reservoirs of antibiotic resistance genes among both pathogenic and nonpathogenic bacteria. Ideally, antibiotics should deactivate rapidly once released into the environment; however, none of the currently used antibiotics meet this criterion. In a study by Zheng et al., a research group from Texas Tech University demonstrated a new antibiotic design with a built-in “OFF” switch responsive to natural stimuli. The design is based on the self-assembly of nanoparticles from environmentally benign building blocks of polymer-grafted cellulose backbones. The group used SAXS performed with SSRL’s beam line 4-2 to probe the interaction of such nanostructures with membranes mimicking mammalian and bacterial cells, and combined it with biochemical assays that established antibiotics cytotoxicity and enzymatic degradation. In their nanostructured forms, the particles are harmless toward mammalian cells, but potent agents against both Gram-positive and Gram-negative bacteria, including clinical multidrug-resistant strains. Upon discharge into the environment, cellulases that are abundant in natural habitats, but not in the human body, shred the particles into antimicrobially-inactive pieces. This study demonstrates mitigation of the environmental footprints of antibiotics using rationally-designed nanoantibiotics that can be dismantled and disabled by biorthogonal chemistry occurring exclusively in natural habitats. | |||
| 11/02/2020 | Mechanistic Insights into Cu(I) Biogenesis of Galactose Oxidase | Structural Molecular Biology Resource | X-ray Absorption and Emission Spectroscopy | Stanford Synchrotron Radiation Lightsource | Chemical and Elemental Information | Together with characterization of the model data, Kβ XES provides a unique opportunity to probe the interaction between Cu(I) and O for O2 activation that is not accessible with other spectroscopic techniques. In this regard, Kβ XES is expected to give important insight into the reactivity of all dioxygen activating Cu(I) metalloprotein systems (i.e., not limited to GO) and to relate the geometric structural assignment (obtained from crystallography or EXAFS methods) to unique electronic properties required for small molecule activation in metalloproteins. | Copper (Cu) active sites in metalloproteins, such as the secretory fungal enzyme galactose oxidase (GO), play essential roles in a wide range of biological processes. One of the most important is O2 activation, which mostly involves Cu(I) active sites. While a variety of spectroscopic methods have been utilized to provide insight into Cu(II) sites, Cu(I) sites are often considered to be “spectroscopically silent” due to their d10 closed-shell nature. A study by Solomon et al. defines the Cu(I) frontier molecular orbital (FMO) that enables this O2 activation and demonstrates the considerable potential of Kβ X-ray emission spectroscopy (XES) in probing the FMO and key bonding interactions in spectroscopically silent Cu(I) active sites. GO catalyzes the oxidation of primary alcohols to aldehydes. Some evidence suggests that GO may also subsequently oxidize the aldehydes to carboxylic acids. This and other research has led to the proposal that the physiological role of GO is to generate hydrogen peroxide (H2O2), perhaps as a defense against pathogenic organisms. GO is also an important component in electrochemical biosensors of galactose that are used for various biotechnology applications. It is therefore critical to understand the mechanism of Cu(I) biogenesis. | Solomon et al. used Kβ XES on SSRL’s beamline 6-2 to characterize a series of Cu(I) compounds with varying biologically relevant ligand systems and compared the results to those of Cu(I) in GO to elucidate the detailed bonding interactions occurring between the metal site and the protein-based ligands. Density functional theory (DFT) calculations were also performed to evaluate the sensitivity of the spectral features to the ligand environment. The study reveals that the Cu(I) site in preprocessed GO (GOpre) is a three-coordinate environment with a 1Tyr/2His structural model. This is consistent with previously reported EXAFS measurements by the same research group. | |
| 09/01/2020 | Tracking Foliar Zinc Absorption in Apple Trees | Structural Molecular Biology Resource | X-ray Fluorescence Imaging | Advanced Photon Source, Stanford Synchrotron Radiation Lightsource | Chemical and Elemental Information | Zinc deficiency in the soil can affect plant health and lead to poor yields, especially in fruit crops. Zinc is a vital micronutrient involved in various cellular processes and it plays a critical role in maintaining the structure of various plant proteins, including transcription factors. Therefore, an interruption or decline in zinc supply substantially affects vegetative development and reproductive success in subsequent years. Zinc fertilization is standard agricultural practice for fruit trees, but achieving successful mitigation can be challenging, particularly in alkaline soils characterized by strong zinc fixation capacity. Nutrient supplementation usually requires application of large quantities of fertilizer, which is cost prohibitive. It is also frequently ineffective for fruit tree crops because the root systems penetrate deep into soil layers where zinc mobility is often poor. In contrast to soil amendments, foliar nutrient application guarantees rapid and targeted uptake via direct delivery to plant tissues during vital growth stages. Foliar nutrient penetration is a complex process that relies on plant leaf surface characteristics, the physicochemical properties of chemical nutrients, supplement type and concentration, and environmental conditions. In a study using synchrotron-based X-ray fluorescence microscopy (XRF) on SSRL’s beamline 2-3, Tian et al. tracked in vivo localization of zinc after foliar fertilizer treatment of apple plants (Malus domestica Borkh.). Apple was selected because of its high sensitivity to zinc deficiency and the vulnerability of apple orchards in areas with calcareous and salt-affected soils, resulting in significant yield losses and fruit quality deterioration. The researchers found that foliar-applied zinc absorption was largely dependent on plant leaf surface characteristics. In particular, abaxial leaf surfaces—the undersides of leaves—absorbed significantly greater zinc concentrations than adaxial leaf surfaces. High-resolution elemental maps revealed that the cell wall’s high binding capacity for zinc contributed to its limited penetration across epidermal cells. Trichome density and stomatal aperture had opposite effects on zinc fertilizer penetration: a relatively high density of trichomes increased the hydrophobicity of leaves, whereas the presence of stomata facilitated foliar zinc penetration. The results indicate that the extent of zinc mobilization is a key factor in its foliar uptake. Low levels of zinc promoted the accumulation of other mineral elements in treated leaves, and the complexation of zinc with phytic acid likely occurred due to exposure to high-zinc conditions. The study provides direct visual evidence for zinc penetration process across the leaf surface, which is important for the development of strategies for zinc biofortification in fruit-bearing crops. | |||
| 01/01/2021 | Soil Amendments and Flooding Cycle Impact Cadmium Bioavailability in Paddy Soil | Structural Molecular Biology Resource | X-ray Absorption and Emission Spectroscopy | Stanford Synchrotron Radiation Lightsource | Chemical and Elemental Information | Chronic cadmium (Cd) exposure is directly associated with osteoporosis, renal dysfunction, and various forms of cancers. In recent years, human activities such as fuel combustion, mining, and industrial manufacturing have caused Cd contamination in soils. Today, consumption of plant-derived foods accounts for approximately 90% of human Cd exposure for the general non-smoking population, with rice being the major dietary source. Paddy rice is grown under episodic cycles of flooding and draining. Typically, paddy water is drained during the later phase of grain filling, resulting in the release of insoluble or sorbed Cd into the soil solution. An estimated 80% of Cd accumulation in rice grain occurs during the grain filling period when paddy water is drained. It is important to understand how Cd is released from the soil solid phase during the drainage period. The effect of the flooding-draining cycles on soil Cd chemical speciation has been previously studied, but a direct correlation between Cd chemical speciation and Cd release kinetics has not been developed. The latter is particularly important for assessing the availability of Cd to uptake by rice plants. Researchers used X-ray absorption spectroscopy on Stanford Synchrotron Radiation Lightsource beamline BL7-3 along with stirred-flow kinetics to investigate the effects of flooding-draining cycles, and CaCO3 and CaSO4 soil amendments, on Cd speciation and release kinetics from a Cd-spiked paddy soil. Extended X-ray absorption fine structure (EXAFS) analysis showed that Cd was predominantly bound to non-iron clay minerals (e.g., Cd-kaolinite, Cd-illite, and Cd-montmorillonite) in the air-dried soil and 1- or 7-day flooded samples. After prolonged flooding (30 and 120 days), Cd-iron mineral complexes became the predominant species. Stirred-flow kinetic analysis showed that both prolonged flooding and the amendments with CaCO3 and CaSO4 decreased the maximum amount and the rate coefficient of Cd release. However, the effect of prolonged flooding was reversed after a short period of draining, indicating that although Cd was immobilized during flooding, it became mobile rapidly after the soil was drained, possibly due to pH decrease and rapid oxidation of CdS. The effects of the amendments on Cd uptake in rice plants were tested in a pot experiment using the same paddy soil without Cd spiking. Data show that amendment with CaCO3, and to a lesser extent CaSO4, decreased the Cd accumulation in two rice cultivars. The combination of CaCO3 amendment and a low-Cd-accumulating cultivar was effective at limiting grain Cd concentration to a low, toxicologically-acceptable level. | |||
| 08/18/2020 | Visualizing Enzymes in Action | Structural Molecular Biology Resource | X-ray Macromolecular Crystallography | Stanford Synchrotron Radiation Lightsource | Molecular Structure | The enzyme 3-Hydroxyanthranilate-3,4-dioxygenase (HAO) is critical for biosynthesis of the coenzyme nicotinamide adenine dinucleotide (NAD+), which in turn is required for biosynthesis in all kingdoms of life. Although the role of this enzyme has long been known, its mechanism and regulation have remained a mystery. HAO is difficult to study because it creates unstable intermediates that can easily form cyclic by-products. Scientists from the University of Texas at San Antonio and the University of Pennsylvania, Philadelphia, have determined the mechanism of the HAO enzyme by studying its crystalline state in which the reaction rate is slowed to minutes. They combined single-crystal UV-Vis spectroscopy and X-ray crystallography to probe changes in the active site and the overall conformation of HAO. The enzyme was first crystallized and then incubated with its aromatic substrate under anaerobic conditions. Subsequently, crystals of enzyme-substrate complex were exposed to oxygen for different lengths of time to interrogate any possible intermediates. The accumulation of catalytic intermediates at various timepoints were identified using the integrated single-crystal UV-Vis spectroscopy system at the Stanford Synchrotron Radiation Lightsource beamline 9-2 and X-ray diffraction data were collected for each corresponding timepoint. The structural analysis provided a step-by-step visualization of the HAO cycle, revealing the intermediates involved and how quinolinic acid production is fine-tuned by the enzyme. | |||
| 01/22/2020 | Efficient and Economical Strategy for Functional Metalloprotein Design | Structural Molecular Biology Resource | Cryo-Electron Microscopy, X-ray Macromolecular Crystallography | Stanford Synchrotron Radiation Lightsource | Molecular Structure | Many proteins exist naturally as symmetrical homo-oligomers or homopolymers. The emergent structural and functional properties of such protein assemblies have inspired extensive efforts in biomolecular design. As synthesized by ribosomes, proteins are inherently asymmetric. Thus, they must acquire multiple surface patches that selectively associate to generate the different symmetry elements needed to form higher-order architectures, a daunting task for protein design. A research group at the University of California San Diego has addressed this problem using an inorganic chemical approach, whereby multiple modes of protein-protein interactions and symmetry are simultaneously achieved by selective, “one-pot” coordination of soft and hard metal ions. The group solved the structures of several synthesized cage variants using data collected on beam line 9-2 at the Stanford Synchrotron Radiation Lightsource. They showed they were able to design interfaces with 2- and 3-fold symmetry axes using cytochrome cb562 variants. By incorporating hydroxamate binding motifs with native chelating residues, they were also able to generate a protein cage with distinct metal ions at the symmetry interfaces. And by varying the ratios of Zn and Fe, they were able to generate different cage symmetries. The new cages closely resembled natural polyhedral protein architectures and are unique in that they are tightly packed and devoid of any large apertures. At the same time, they assemble and disassemble in response to diverse stimuli, owing to their heterobimetallic construction on minimal interprotein-bonding footprints. With varying stoichiometries, these protein cages represent some of the most compositionally complex protein assemblies obtained by design. | |||
| 03/10/2020 | Introducing Positive Charges in an Artificial Enzyme Increases CO2 Hydrogenation | Structural Molecular Biology Resource | X-ray Macromolecular Crystallography | Environmental Molecular Sciences Laboratory, Stanford Synchrotron Radiation Lightsource | Molecular Structure | Metalloenzymes play central roles in many energy transduction reactions by facilitating the storage of energy in chemical bonds until the energy needs to be used. Because metalloenzymes are very efficient, fast, and selective in the conversion and storage of energy under ambient conditions, they are ideal models upon which to base synthetic mimics capable of efficiently interconverting electrical and chemical energies. One of the critical features responsible for the high reactivity and specificity of metalloenzymes is the protein scaffold surrounding the metal at the active site. Protein scaffolds are composed of elements of secondary structure dominated by α-helices and β-sheets which, in turn, fold into higher order tertiary structures with a well-defined spatial arrangement of hydrophobic and hydrophilic residues. These highly evolved three-dimensional spaces tune specific chemical reactions catalyzed by the metal at the active site. A research group at Washington State University has synthesized an artificial enzyme capable of CO2 hydrogenation containing the small molecular complex [Rh(PEt2NglycinePEt2)2]– bound to a protein scaffold. Point mutations were made in the outer coordination spheres for several complexes, producing positive charges at various distances from the Rh center. The structures were solved using data acquired on beam line 12-2 at the Stanford Synchrotron Radiation Lightsource. The structure of a complex with a 3-fold greater turnover rate showed that its positive charges were close (8-9 Å) and oriented towards the Rh site. This work should expand the synthetic toolset for the development of more efficient enzymes. | |||
| 12/09/2019 | Tuning the Catalytic Bias of Hydrogenase Enzymes | Structural Molecular Biology Resource | Synchrotron Infrared Hyperspectral Imaging, X-ray Macromolecular Crystallography | Environmental Molecular Sciences Laboratory, Stanford Synchrotron Radiation Lightsource | Molecular Structure, Chemical and Elemental Information | Hydrogenase enzymes catalyze the seemingly simple interconversion of protons and electrons with hydrogen (H2) gas. These enzymes are ubiquitous in nature, being found in bacteria and archaea as well as in single-celled eukaryotes. They are important in H2 gas cycling and interspecies H2 gas transfer as well as in the energetics of many microbial systems. The unifying feature of all hydrogenases of the [FeFe]-type is a complex and unique metallocluster termed the “H-cluster.” The H-cluster contains a conventional cubane [4Fe-4S] subcluster that is bridged through a cysteinyl thiolate to a unique 2Fe subcluster having carbon monoxide, cyanide, and azadithiolate ligands. A group of researchers at Washington State University, in collaboration with Stanford Synchrotron Radiation Lightsource X-ray macromolecular crystallography (MX) scientists, solved several hydrogenase structures from Clostridium pasteurianum to very high resolution using beam line12-2 and found dual conformations of residues near the H-cluster. X-ray free-electron laser (XFEL) data were also collected at the Linac Coherent Light Source X-ray Pump Probe station, which provided evidence that the different conformations represented different oxidation states. Together with electrochemical, spectroscopic and theoretical calculations, the group found that key amino acids within the H-cluster adopted different conformations as a function of oxidation state. This structural flexibility makes it possible to modulate the properties of the electronic structure of the H-cluster via secondary, noncovalent interactions. The researchers propose that the dynamic changes observed in C. pasteurianum are important for imposing a neutral catalytic bias or an efficacy in catalyzing both H2 oxidation and proton reduction at similar rates. | |||
| 03/08/2018 | Safety Concerns Over Tungsten Accumulation in Bone | Center for BioMolecular Structure | X-ray Fluorescence Imaging | National Synchrotron Light Source II | Chemical and Elemental Information | New research shows how and where tungsten, which has been associated with childhood lymphocytic leukemia, accumulates in the bones of mice exposed to the element through drinking water. X-ray fluorescence spectroscopy showed that tungsten accumulated in bone tissue, with some sites having around 10-fold greater intensities than background levels. The findings raise doubts over the once-universal assumption that tungsten poses little or no health risk to the general human population. | |||
| 07/08/2019 | Nanoparticle Properties Influence Nutrient Transport in Plants | Center for BioMolecular Structure, Structural Molecular Biology Resource | X-ray Absorption and Emission Spectroscopy, X-ray Fluorescence Imaging | National Synchrotron Light Source II, Stanford Synchrotron Radiation Lightsource | Chemical and Elemental Information | Plant nanobiotechnology promises transformative solutions to the most vexing problems threatening global food security, including drought, disease, and soil nutrient deficiencies. However, the lack of cost-effective methods for delivering nanomaterials to the precise locations where they become active, such as inside vascular tissues or organelles, impedes these technological innovations. Scientists used X-ray fluorescence microscopy and X-ray absorption spectroscopy to evaluate the influence of nanoparticle surface charge and differences in root structure and vasculature on cerium distribution within plants. They found that both nanoparticle surface chemistry and plant morphology significantly impacted the uptake and distribution of cerium nanoparticles. The findings not only provide insight into how plant structural features influence nanoparticle behavior but also how surface charge can be tailored for targeted delivery of nutrients to specific plant organs. | |||
| 10/17/2018 | 3.7-Billion-Year-Old Rock Structures Formed by Tectonics, Not Life | Center for BioMolecular Structure | X-ray Fluorescence Imaging | National Synchrotron Light Source II | Chemical and Elemental Information | Scientists reevaluated evidence of life in 3.7-billion-year-old rock structures using improved imaging techniques and determined they were of geological origin, not biological origin as previously thought. The structures had been considered the earliest evidence for life on Earth. The morphology, layering, mineralogy, chemistry, and geological context of the structures were originally attributed to the formation of microbial mats in a shallow marine environment 3.7 billion years ago at the start of Earth’s rock record. Previous studies of the rock samples employed millimeter scale information using laser ablation analysis to determine the origin of the formations. In the more recent study, a team of scientists used micrometer X-ray florescence analysis at the National Synchrotron Light Source II (NSLS-II) to reexamine the rock structures. The Submicron Resolution X-ray (SRX) beamline revealed crucial new evidence based on the elemental make-up of the rock structures. This evidence supports the conclusion that the rock formations are of non-biological origin. | |||
| 09/07/2018 | X-ray Nanofluorescence Tomography of Single Bacterial Cells | Center for BioMolecular Structure | Hard X-ray Tomography, X-ray Fluorescence Imaging, X-ray Ptychography | Advanced Photon Source, National Synchrotron Light Source II | Chemical and Elemental Information | X-ray fluorescence (XRF) nanotomography was used to image elemental distribution in individual E. coli bacterial cells using a sub-15 nm beam at the Hard X-ray Nanoprobe beamline (HXN, 3-ID) at the National Synchrotron Light Source II. The measurements were simultaneously combined with ptychography to image the cells’ structural components. The results showed a generally uniform distribution of calcium but an inhomogeneous zinc distribution, most notably with concentrated regions of zinc at the polar ends of the cells. XRF microscopy is a growing approach for imaging the trace element concentration, distribution, and speciation in biological cells at the nanoscale. Moreover, three-dimensional nanotomography provides the added advantage of imaging subcellular structure and chemical identity in three dimensions without the need for staining or sectioning of cells. The work demonstrates that simultaneous two-dimensional ptychography and XRF nanotomography can be performed with a sub-15 nm beam size on unfrozen biological cells to co-localize elemental distribution and nanostructure simultaneously. This multimodal approach presents new possibilities in understanding subcellular biochemistry in individual organelles, which are usually analyzed at the population level. | |||
| 12/10/2018 | Nanoparticle Surface Chemistry Influences Journey Through Tomato Plant | Center for BioMolecular Structure | X-ray Absorption and Emission Spectroscopy, X-ray Fluorescence Imaging | Advanced Photon Source, National Synchrotron Light Source II | Chemical and Elemental Information | Terrestrial ecosystems are a major sink for manufactured nanomaterials (MNM) released unintentionally or used intentionally in agrochemical formulations. These nanomaterials can be taken up by plants and transferred to herbivores. The surface chemistry of manufactured nanoparticles can have a profound impact on their uptake and translocation in plants. However, there is a limited understanding of the tissue, cellular, and subcellular basis for this. Researchers used a novel hard X-ray nanoprobe with unprecedented spatial resolution (<15 nm) to reveal details about the effects of the surface chemistry of the nanoparticle cerium dioxide (CeO2) on its uptake and translocation in tomato (Solanum lycopersicum). CeO2 is a good model material for studying plant-MNM interactions because it is relatively insoluble in typical plant growth media and can be easily measured and tracked by different imaging and spectroscopic methods. CeO2 MNM are among the most widely used nanomaterials found in fuel additives, polishing agents, industrial catalysts, and have recently received interest for use as plant growth promoters in agricultural production. This study provides critical information on how particle surface chemistry influences the biodistribution and cellular localization of nanomaterials in plants using high resolution X-ray imaging of nanomaterials in plant cells. This information enhances the ability to predict how nanomaterial properties influence the uptake, transformations, and subsequent trophic transfer of nanomaterials in terrestrial food webs. | |||
| 11/07/2017 | Characterization of Elusive Yeast Protein Complex Involved in Autophagy | Center for BioMolecular Structure | Solution X-ray Scattering | National Synchrotron Light Source II | Molecular Structure | Autophagy is a form of cellular self-cleaning, where cytoplasmic material destined for degradation is captured in double-membrane compartments that mature and fuse with a degradative organelle. Autophagy is essential to maintain cellular homeostasis, respond to cellular stress, and prevent the accumulation of material that could damage the cell. Highly dynamic proteins engaged in autophagy are very challenging to study using standard techniques that reveal a 3D structure. Autophagy is initiated by the Atg1 protein complex. Whereas recent work has provided functional and mechanistic insight into many components of the Atg1 complex, one member of this complex—Atg20—has remained relatively uncharacterized. Scientists used small-angle X-ray scattering (SAXS) at the LiX beamline at NSLS-II to provide the first insight into the structure and function of Atg20 in Saccharomyces cerevisiae yeast, including identifying an amphipathic helix required for efficient autophagy and membrane tubulation. | |||
| 03/24/2020 | Critical Structural Differences Between Seasonal and Pandemic Influenza | Center for BioMolecular Structure | Solution X-ray Scattering, X-ray Macromolecular Crystallography | National Synchrotron Light Source II | Molecular Structure | Among all influenza pandemics, the 1918 influenza is considered the worst in human history. To investigate differences in virulence among influenza strains, researchers studied the nonstructural protein 1 (NS1) of the 1918 influenza strain and compared it to a seasonal strain. NS1 is an important factor influencing influenza infection severity because it antagonizes host defense mechanisms. The researchers focused on NS1’s interaction with the human phosphoinositide 3-kinase (PI3K), to which NS1 binds through its p85β subunit, and presented the mechanism underlying the molecular recognition of p85β by NS1. They found that the structure of 1918 NS1 is highly dynamic, whereas NS1 of a seasonal influenza strain is mostly static. Moreover, the respective NS1 proteins bind to p85β with drastically different binding affinities and kinetics. The findings provide a mechanistic insight into strain-dependent behaviors of NS1 proteins, which remains elusive despite its importance in understanding the virulence of influenza viruses. | |||
| 03/24/2020 | Tunable Molecular Coating Stabilizes "DNA Origami" Structures | Center for BioMolecular Structure | Solution X-ray Scattering | National Synchrotron Light Source II | Molecular Structure | DNA nanotechnology provides a structural toolkit for the fabrication of programmable DNA nano-constructs through the folding of long, flexible DNA chains into desired shapes at the nanoscale. However, the use of such nano-constructs, known as “DNA origami” structures, in biomedical applications is challenging due their limited structural integrity in complex biological fluids. Researchers report a class of tailorable molecular coatings, called peptoids, which can efficiently stabilize three-dimensional wireframed DNA constructs under a variety of biomedically relevant conditions, including magnesium-ion depletion and presence of degrading nuclease. The peptoid-coated DNA constructs offered a controllable anticancer drug release and an ability to display functional biomolecules on the DNA surfaces. The study demonstrates an approach for building multifunctional and environmentally robust DNA-based molecular structures for nanomedicine and biosensing. | |||
| 01/10/2018 | Novel Architecture of a Bacterial Plasma Membrane Transporter | Center for BioMolecular Structure | X-ray Macromolecular Crystallography | National Synchrotron Light Source II | Molecular Structure | Researchers used X-ray diffraction to reveal a crystal structure for an O-antigen transporter, called Wzm-Wzt, in an unexpected channel-forming conformation. O-antigens are cell surface polysaccharides of many Gram-negative bacterial pathogens that aid in escaping the host’s innate immune responses. A widespread O-antigen biosynthesis mechanism involves the synthesis of the lipid-anchored polymer on the cytosolic face of the inner membrane, followed by transport to the periplasmic side where it is ligated to the lipid A core to complete a lipopolysaccharide molecule. The channel-forming structure stands in contrast to the classical “altering access” mechanism typically used by this class of transporters. Such transmembrane channels are usually confined to ion channels and various toxins and are not observed in transporters that move solutes against their concentration gradient. However, the particular length of O-antigens requires the Wzm-Wzt transporter to utilize this special transport mechanism. | |||
| 11/16/2017 | 3D Protein Structure Aids Search for Vaccine | Center for BioMolecular Structure | X-ray Macromolecular Crystallography | National Synchrotron Light Source II | Molecular Structure | Human metapneumovirus (hMPV) is a frequent cause of bronchitis in young children, but vaccine development has been hindered by an inability to produce the hMPV F glycoprotein, which mediates virus–cell membrane fusion and is the primary target of neutralizing antibodies. Using cryogenic macromolecular crystallography and modeling based on the known structure of a homologous molecule, scientists revealed the 3D structure of the hMPV F glycoprotein. This new knowledge about the viral structure should facilitate development of effective hMPV vaccine candidates. | |||
| 08/09/2018 | Activation Mechanism of a Protein Kinase Reveals Role in Cancer Proliferation | Center for BioMolecular Structure | X-ray Macromolecular Crystallography | National Synchrotron Light Source II | Molecular Structure | Akt 1 protein kinase plays an essential role in cancer proliferation, which defines cancer aggressiveness. Understanding the full functions of the protein could help advance cancer therapies. Scientists revealed a different activation mechanism of Akt1 protein kinase by closely studying its molecular interactions. The results provide a new framework for understanding how Akt 1 is controlled in cell signaling and suggest distinct functions for differentially modified Akt forms. Understanding the full functions of the protein could help advance cancer therapies. | |||
| 10/08/2018 | Searching for New Ways to Fight Bacterial Resistance | Center for BioMolecular Structure | X-ray Macromolecular Crystallography | National Synchrotron Light Source II | Molecular Structure | Natural rifamycin antibiotics and their synthetic derivatives (Rifs) target bacterial RNA polymerases (RNAPs) and are widely used to treat infections, including tuberculosis. The utility of these compounds is threatened by increasing bacterial resistance to the drugs (RifR). As resistance mechanisms found in clinical settings may also occur in natural environments, researchers postulated that bacteria could have evolved to produce rifamycin congeners active against clinically relevant resistance phenotypes. A survey of soil metagenomes identified a tailoring enzyme-rich family of gene clusters encoding biosynthesis of rifamycin congeners called kanglemycins (Kangs) that display potent in vivo and in vitro activity against the most common clinically relevant RifR mutations. Structural and mechanistic analyses revealed the basis for Kang inhibition of RifR RNAP. Unlike Rifs, Kangs function through a mechanism that includes interfering with 5′-initiating substrate binding. The results suggest that examining soil microbiomes for new analogues of clinically used antibiotics may uncover metabolites capable of circumventing clinically important resistance mechanisms. | |||
| 06/21/2019 | Light Teaches Coenzymes New Tricks | Center for BioMolecular Structure | X-ray Macromolecular Crystallography | National Synchrotron Light Source II | Molecular Structure | During synthesis of organic molecules, light is widely used to excite electrons in a substrate or catalyst, opening up reactive pathways to a desired product. In biological systems, light is used sparingly in this way, but coenzymes such as flavins can be artificially driven to excited states by light. Photoexcitation of promiscuous flavoenzymes can thus furnish previously unknown biocatalytic reactions. Researchers investigated this reactivity of coenzymes and found a suite of flavoenzymes from the bacterium Gluconobacter oxydans that catalyze asymmetric radical cyclization when exposed to light. X-ray macromolecular crystallography revealed that photoexcitation enabled the flavin-dependent “ene”-reductases to convert starting materials containing an α-chloroamide and an alkene into five-, six-, seven-, or eight-membered lactams. After formation of a prochiral radical, the enzyme guides the delivery of a hydrogen atom from flavin—a challenging feat for small-molecule chemical reagents. The initial electron transfer occurs through direct excitation of an electron donor-acceptor complex that forms between the substrate and the reduced flavin cofactor within the enzyme active site. *Image Use: Readers may view, browse, and/or download this image for temporary copying purposes only, provided these uses are for noncommercial personal purposes. Except as provided by law, this image may not be further reproduced, distributed, transmitted, modified, adapted, performed, displayed, published, or sold in whole or in part, without prior written permission from the publisher [AAAS]. | |||
| 05/21/2019 | Structural Basis of Bacterial Spore Germination | Center for BioMolecular Structure | X-ray Macromolecular Crystallography | National Synchrotron Light Source II | Molecular Structure | Bacterial endospores are found throughout the environment, with some acting as vectors of food spoilage, food poisoning, and infectious diseases. Promoting efficient spore germination could increase spore sensitivity to inactivating measures. Germination of Bacillus spp. bacteria spores is induced by the interaction of specific nutrient molecules with germinant receptors localized in a spore’s inner membrane. Germinant receptors typically consist of three subunits, referred to as A, B, and C. Researchers used X-ray macromolecular crystallography to determine the crystal structure of the N-terminal domain (NTD) of the A subunit of the Bacillus megaterium germinant receptor, GerK3. Crystal structure, molecular docking, and biophysical and genetic analyses indicate the presence of a binding pocket at the interface between the two subdomains in the NTD of the A subunit that accommodates specific germinant molecules and plays a critical role in spore germination. The results can be explored for the development of germinant analogues as either potentiators or inhibitors of spore germination. | |||
| 05/07/2020 | Mechanism of Activation of a Plant Metacaspase | Center for BioMolecular Structure | X-ray Macromolecular Crystallography | National Synchrotron Light Source II | Molecular Structure | Plant metacaspase enzymes mediate many important cellular functions, including programmed cell death, stress and immune responses, and resistance to pathogens. Using X-ray macromolecular crystallography, researchers determined the crystal structures comprising Metacaspase 4 from Arabidopsis thaliana (AtMC4) and characterized its dependence on Ca2+ for activation. Understanding the mechanisms of metacaspase-mediated responses offers a basis for future bioengineering to enable the design of more sustainable crops and biofuels. | |||
| 04/14/2021 | Molecular Fragment Screening Identifies Potential SARS-CoV-2 Inhibitors | Center for BioMolecular Structure, Structurally Integrated Biology for the Life Sciences, Structural Molecular Biology Resource | X-ray Macromolecular Crystallography | Advanced Light Source, National Synchrotron Light Source II, Stanford Synchrotron Radiation Lightsource | Molecular Structure | Research published in Science Advances provides a template for developing direct-acting antiviral drugs, with novel modes of action, to combat COVID-19 by suppressing the SARS-CoV-2 viral infection.  Surface representation of a SARS-CoV-2 macrodomain (Nsp3 Mac1) bound to the host signaling molecule ADPr (cyan), which inhibits the host immune response. [Credit: M. Schuller, et al. 2021. DOI:10.1126/sciadv.abf8711] A large-scale crystallographic screening effort combined with computational docking identified 234 new molecules capable of targeting the active site of Mac1. These fragment structures provide starting points for developing potent SARS-CoV-2 Mac1 inhibitors. | |||
| 06/13/2018 | Structure of a Nuclear Membrane Gatekeeper | Center for BioMolecular Structure | X-ray Macromolecular Crystallography | National Synchrotron Light Source II | Molecular Structure | Nuclear pore complexes (NPC) are macromolecular machines which perforate the nuclear envelope and control passage of molecules such as mRNA from the cell’s nucleus to the cytoplasm. But how NPCs directly participate in macromolecular transport has been poorly understood. A team of scientists crystalized a protein complex which partially comprises NPCs—Gle1•Nup42—from three different organisms, including humans. Next, they used X-rays from the Frontier Microfocusing Macromolecular Crystallography (FMX) beamline to reveal the 3-D crystal structure for all three organisms, uncovering the evolutionarily conserved binding mode for each. The 3-D structure of the protein complex in humans showed how it ensures mRNA transport. Lastly, they found that Gle1 mutations associated with motor neuron diseases possess severe thermostability defects, suggesting that nucleoporin misfolding contributes to disease. The results provide the foundation for further mechanistic analyses of mRNA export in humans. | |||
| 09/23/2019 | Mechanism of Regulatory Protein's Dual Role as Inhibitor and Activator | Center for BioMolecular Structure | X-ray Macromolecular Crystallography | National Synchrotron Light Source II, Stanford Synchrotron Radiation Lightsource | Molecular Structure | Researchers combined genetic, cellular, molecular, and X-ray crystallographic techniques to understand how SDS22, an evolutionarily highly conserved regulatory protein, interacts with the protein phosphatase 1 (PP1) enzyme and influences its activity. PP1 accounts for more than half of dephosphorylation reactions in eukaryotes and is exceptionally well conserved from fungi to humans in both sequence and function. PP1 is involved in numerous fundamental cellular processes including mitosis, protein synthesis, and regulation of membrane receptors and channels. PP1 is controlled by more than 200 regulatory proteins which enable it to bind to a wide range of substrates in a highly specific manner. SDS22 is known to act as both an inhibitor and activator of PP1, but the mechanism used to achieve this dual role has been unclear. X-ray macromolecular crystallography revealed that SDS22 exclusively binds and traps a previously undescribed metal-deficient conformation of PP1, rendering it inactive. Once the complex forms, it does not permanently dissociate until PP1 binds the missing metal ion. Thus, SDS22 acts as a “PP1 trap,” providing a pool of PP1 poised for the rapid formation of new holoenzymes in response to dynamically changing cellular events. Taken together, the data reveal the critical role of metal binding in PP1 activation and provide fundamental insights into the mechanisms by which SDS22 inhibits and stabilizes PP1 prior to holoenzyme formation. Understanding the mechanisms of PP1 holoenzyme assembly and activation could lead to novel therapies for diseases associated with PP1 signaling. | |||
| 04/02/2013 | Major Scaffold Component in Nuclear Pore Complex | Structural Molecular Biology Resource | Solution X-ray Scattering | Stanford Synchrotron Radiation Lightsource | Molecular Structure | Nuclear pore complexes (NPC) are large, octagonally symmetric, cylindrical macromolecular assemblies formed at the fusion of the inner and outer nuclear envelope membranes in eukaryotes. The NPC is responsible for the rapid, active, and selective exchange of macromolecules between the nucleus and cytoplasm. As both the gatekeepers of nucleocytoplasmic transport and a platform for the organization of numerous nuclear activities, NPCs contribute to the regulation of myriad physiological processes, including gene expression, cell cycle control, and spindle and kinetochore assembly. NPCs are composed of proteins termed nucleoporins (Nups), forming an annular structure composed of the nuclear ring, cytoplasmic ring, a membrane ring, and two inner rings. Nup192 is a major component of the NPC’s inner ring. Small angle X-ray scattering (solution X-ray scattering) and electron microscopy (EM) studies determined the crystal structure of Saccharomyces cerevisiae Nup192 residues 2–960 (ScNup192(2–960)), and revealed that ScNup192(2–960) can undergo long-range transition between “open” and “closed” conformations. Researchers then obtained a structural model of full-length ScNup192 based on EM, the structure of ScNup192(2–960), and homology modeling. Evolutionary analyses using the ScNup192(2–960) structure suggest that NPCs and vesicle-coating complexes are descended from a common membrane-coating ancestral complex. Finally, experimental suppression of Nup192 expression led to compromised nuclear transport, suggesting a possible role for Nup192 in modulating the permeability of the NPC central channel and ensuring efficient nuclear transport. | |||
| 05/24/2012 | Modulation of Plant Hormones by GH3 Proteins | Structural Biology Center | X-ray Macromolecular Crystallography | Advanced Photon Source | Molecular Structure | Plants respond to developmental cues and environmental stresses by controlling the level and activity of various hormones. Within the auxin-responsive GH3 enzyme family, a highly adaptable protein scaffold enables the evolution of promiscuous activity and leads to the diversification of substrate specificity and the evolution of metabolic control systems. Newly reported crystal structures for two acyl acid amido synthetase enzymes belonging to the GH3 family from Arabidopsis provide a glimpse into substrate recognition and the control of hormones involved in plant growth, development, and defense. The findings lend insight into the reaction chemistries that add functional diversity to hormone signaling pathways. | |||
| 10/29/2013 | Common Physical Processes in Different Biomass Pretreatments | Center for Structural Molecular Biology | Small-Angle Neutron Scattering, X-ray Macromolecular Crystallography | Spallation Neutron Source/High Flux Isotope Reactor | Molecular Structure | Lignocellulosic biomass, a potentially important renewable source of energy and chemical feedstocks, resists degradation to glucose during industrial hydrolysis processes and thus requires expensive thermochemical pretreatments. Understanding the mechanism of biomass breakdown during these pretreatments can lead to more efficient use of biomass. Researchers used X-ray fiber diffraction and small-angle neutron scattering (SANS) at the CG-2 instrument at the High Flux Isotope Reactor (HFIR) facility of Oak Ridge National Laboratory to probe the molecular structure of biomass at different length scales during steam explosion pretreatment. Next, they used the data in molecular dynamics simulations to reveal two fundamental processes responsible for the resulting morphological changes: cellulose dehydration and lignin-hemicellulose phase separation. They further found that the basic driving forces occurring during steam explosion are the same as in other leading thermochemical pretreatments, such as dilute acid pretreatment and ammonia fiber expansion. The findings suggest that new pretreatments and plant modifications that promote lignin and hemicellulose phase separation and increase the porosity of the cell wall matrix while preventing increases in cellulose crystallization as a result of dehydration can improve biomass conversion. | |||
| 07/17/2013 | Lessening X-ray Damage Speeds Protein Discovery | Structural Biology Center | X-ray Macromolecular Crystallography | Advanced Photon Source | Molecular Structure | Synchrotron facilities like the DOE’s Advanced Photon Source (APS) at Argonne contain particle accelerator systems designed to produce extraordinarily bright and high-energy X-ray beams. At these facilities, scientists can peer deeply into the atomic structure of molecules using the method of X-ray crystallography. However, the use of powerful X-ray beams to study protein crystals poses a dilemma: the beams provide the best tool for understanding a protein’s structure and biological function, but they often damage the crystal. Scientists examined three different X-ray-based methods for solving protein structures and recommended one called “submicrometer line focusing” as the most promising for easing the dilemma. The beam strikes the protein crystal with an area smaller than a micrometer, thereby minimizing the impact area. The beam is also focused as a vertical line, delivering a more concentrated dose of X-rays per area. The researchers also suggested using a new lens they designed that breaks the powerful beam into many mini-beams, spaced far enough apart that the damage one mini-beam creates lies outside the area probed by neighboring mini-beams. | |||
| 02/07/2013 | Lipid Bilayer Composition Regulates Function of Lipid Rafts in Model Cell Membranes | Center for Structural Molecular Biology | Small-Angle Neutron Scattering | Spallation Neutron Source/High Flux Isotope Reactor | Molecular Structure | Floating freely in cell membranes, lipid rafts are organizing centers for membrane-mediated processes. Neutron scattering techniques were used to characterize lipid domain size transitions from nanometers to micrometers in a four-component biomimetic lipid mixture. Results suggest that reversible changes in lipid composition may regulate the size of functional domains in lipid rafts. The BioSANS instrument at the ORNL Center for Structural Molecular Biology and the EQ-SANS instrument (BER) at the ORNL Spallation Neutron Source (BES) were used in this study. | |||
| 12/09/2012 | Watching the Evolution of a Protein’s Function | Structural Molecular Biology Resource | X-ray Absorption and Emission Spectroscopy | Stanford Synchrotron Radiation Lightsource | Chemical and Elemental Information | Using directed evolution—a process by which organisms or genes are artificially evolved to develop nucleic acid mutations or produce new proteins—researchers have determined the structural basis for converting a noncatalytic small protein into an effective enzyme for linking RNA molecules. | A major challenge to large-scale biofuels production is developing enzymes that are highly efficient at converting biomass components into usable fuels. | Enzymes are proteins structurally configured to catalyze conversions of substrates to products. Thousands of protein structures are known, including those of many valuable enzymes. However, much less is known about how small changes in a protein’s composition can alter its three-dimensional structure, control its catalytic efficiency, or even convert a protein with no catalytic function into an efficient catalyst. Scientists in this study used an extended X-ray absorption fine structure (EXAFS) station at the Stanford Synchrotron Radiation Lightsource to determine the active-site structure of a newly synthesized enzyme. The EXAFS experiments showed the exact chemical environment of each zinc atom in the new enzyme, leading to an explanation of why it developed the catalytic activity. | |
| 08/15/2013 | Multifunctionality of an Ebola Virus Protein | Structural Biology Center, Structural Molecular Biology Resource | X-ray Macromolecular Crystallography | Advanced Light Source, Advanced Photon Source, Stanford Synchrotron Radiation Lightsource | Molecular Structure | Proteins, particularly viral proteins, can be multifunctional, but the mechanisms behind multifunctionality are not fully understood. Scientists illustrated through multiple crystal structures, biochemistry, and cellular microscopy that the Ebola virus protein VP40 rearranges into different structures, each with a distinct function required for the ebolavirus life cycle. A butterfly-shaped VP40 dimer traffics to the cellular membrane. Once there, electrostatic interactions trigger rearrangement of the polypeptide into a linear hexamer. These hexamers construct a multilayered, filamentous matrix structure that is critical for budding and resembles tomograms of authentic virions. A third structure of VP40, formed by a different rearrangement, is not involved in virus assembly but instead uniquely binds RNA to regulate viral transcription inside infected cells. These results provide a functional model for ebolavirus matrix assembly and the other roles of VP40 in the virus life cycle and demonstrate how a single wild-type, unmodified polypeptide can assemble into different structures for different functions. | |||
| 09/26/2014 | Understanding Nitrogen Fixation in Bacteria | Structural Molecular Biology Resource | X-ray Macromolecular Crystallography | Stanford Synchrotron Radiation Lightsource | Molecular Structure | Living organisms depend on nitrogen fixation to convert atmospheric nitrogen (N2) to a form they can incorporate into the basic building blocks of life, such as DNA and amino acids. Nitrogenase is the only known enzyme capable of fixing nitrogen. Therefore, understanding how nitrogenase performs its multi-electron reduction of N2 is highly important in ammonia fertilizer production, for energy efficiency (because industrial ammonia production consumes enormous amounts of energy), and for addressing climate change. Scientists used X-ray macromolecular crystallography to determine the structure of the MoFe protein, one of two metalloproteins that comprise nitrogenase in the soil bacterium Azotobacter vinelandii, and how substrates bind to the enzyme’s active site. The structure and conformational changes that occur when bound to a substrate provide insight into how the N2 triple bond is reduced. | |||
| 11/10/2014 | Self-Assembling Porous Protein Cage Designed for Potential Biotech Applications | Structurally Integrated Biology for the Life Sciences | Solution X-ray Scattering, X-ray Macromolecular Crystallography | Advanced Light Source | Molecular Structure | Scientists used small-angle X-ray scattering (SAXS; aka, solution X-ray scattering) to efficiently and quickly visualize the self-assembly process of a designed cage-like protein in different conditions. The study demonstrates that accurate design of large porous protein assemblies with specific shapes is feasible, and that specificity improvements may require limiting flexibility to select against alternative forms. The hollow, cube-shaped protein cage could potentially deliver proteins or other chemicals to specific locations for medical, energy, and other applications. These results provide a foundation for the design of advanced materials with applications in bio-nanotechnology, nanomedicine, and material sciences. | |||
| 08/25/2015 | Modular Construction on a Biomolecular Scale | Structural Molecular Biology Resource | X-ray Macromolecular Crystallography | Stanford Synchrotron Radiation Lightsource | Molecular Structure | The spherical protein ferritin was used to synthesize a porous 3-D crystalline framework material, and further engineered to have metal hubs on its surface. Organic molecules bridge these hubs, and in a controlled way create a porous material with potential uses from efficient fuel storage to carbon capture and conversion. The study shows the great potential for proteins as building blocks with exquisite properties of electron transfer and bioinspired catalysis. | |||
| 06/10/2015 | Archaeal Enzyme Plays Key Role in Atmospheric Carbon Turnover | Structural Biology Center | X-ray Macromolecular Crystallography | Advanced Photon Source | Molecular Structure | Researchers from the U.S. Department of Energy’s Argonne National Laboratory and the University of Tennessee found that microorganisms called archaea living in marine sediments use completely novel enzymes to break down organic matter into carbon dioxide. Scientists are uncertain how fast archaea process carbon and whether the release is accelerating. Once researchers have these statistics, they may find ways to better predict the environment’s response to a changing climate. This understanding starts at the molecular level. Using resources at the Advanced Photon Source, a DOE Office of Science User Facility, and the Advanced Protein Characterization Facility, the research team isolated and crystallized bathyaminopeptidase (BAP), one of the enzymes found in the archaea, to look into its structure and observe how it operates. They found that BAP plays an important role in breaking down proteins and, consequently, the turnover of atmospheric carbon. | |||
| 09/08/2015 | Manganese Reduction-Oxidation Drives Plant Debris Decomposition | Berkeley Synchrotron Infrared Structural Biology Imaging Program, Structural Molecular Biology Resource | Synchrotron Infrared Hyperspectral Imaging, X-ray Absorption and Emission Spectroscopy, X-ray Fluorescence Imaging | Advanced Light Source, Stanford Synchrotron Radiation Lightsource | Chemical and Elemental Information | Microbial decomposition of plant debris (litter) regulates nutrient availability, ecosystem productivity, and carbon (C) cycling. Historically, scientists thought climate—primarily temperature and precipitation—regulated the rate of litter decomposition, which then influenced the rate at which nutrients become available and C contained in the litter was released back into the atmosphere as the greenhouse gas carbon dioxide (CO2). Recently, however, evidence has shown that geochemical factors also influence litter decomposition rates. A research team has shown that long-term litter decomposition rates in forest ecosystems are closely related to the process of manganese (Mn) reduction-oxidation (redox) in a variety of forest ecosystems. A strong correlation had been observed previously between litter Mn content and decomposition rates, but the underlying mechanisms were not well understood. In redox cycling, redox governs Mn reactions, frequently mediated by microbes, which accumulate and oxidize Mn in the litter, then using the oxidized Mn species to break down plant cells. By pairing high-resolution chemical imaging analysis with a long-term litter decomposition experiment under field conditions, the researchers discovered that litter decomposition is tightly coupled to redox cycling of Mn. The researchers discovered why Mn exerts such a strong control on litter decomposition rates and the mechanism of the underlying biogeochemical process, observing molecular changes within the litter as it decomposed over the years and probing structural changes within the very large biomolecules present in litter that frequently are not directly detectable by other means. This tunable technique also allowed the scientists to selectively ionize different biopolymers present in the litter, such as lignin, which is important because it lends rigidity to plant cell walls and protects litter from microbial decay. In microenvironments within the litter, microbes actively cycle Mn as they colonize and break down the litter, and research indicates that biogeochemical constraints on bioavailability, mobility, and Mn reactivity in the plant-soil system may have a profound impact on litter decomposition rates. Understanding more about the mobility and reactivity of Mn in the plant-soil system also helps researchers improve their ability to accurately predict carbon cycling trends in ecosystems, contributing to greater insights into climate change patterns. Instruments and FacilitiesBeamlines 10.3.2 (X-ray microprobe capabilities) and 1.4.3 (Fourier transform infrared spectromicroscopy) at Advanced Light Source (ALS) at Lawrence Berkeley National Laboratory; beamline 4-3 at Stanford Synchrotron Radiation Lightsource (SSRL). | |||
| 12/07/2016 | Using Soft X-ray Ptychography to Investigate a Bacterium’s Inner Compass | National Center for X-Ray Tomography | X-ray Absorption and Emission Spectroscopy, X-ray Ptychography | Advanced Light Source | Chemical and Elemental Information | Magnetotactic bacteria (MTB) synthesize chains of magnetic nanocrystals (magnetosomes) that interact with the Earth’s magnetic field like an inner compass needle, simplifying their search for optimum environments. Some studies investigating how these magnetosomes form suggest that hematite or amorphous ferrihydrite act as precursors, while others posit that they are formed directly from solution-phase Fe(II) and Fe(III). Thus, identifying the chemical states of precursors would be a good way to differentiate among competing models. Ptychography is a coherent diffractive imaging technique with high resolution (7 nm in this work) and excellent chemical-state sensitivity. Researchers obtained ptychographic absorption and phase spectra of magnetosomes from a marine MTB (Magnetovibrio blakemorei strain MV-1). The data, obtained from mature magnetosomes, immature magnetosomes, precursor regions, and the gaps between magnetosome chains, indicated that different iron species can coexist in a single cell. Based on the results, the researchers proposed a model in which soluble Fe(II) is taken up from the environment and partially oxidized to Fe(III), which in turn is then oxidized and transformed into hematite and ultimately into magnetite. Researchers also used ptychography to measure the X-ray magnetic circular dichroism (XMCD) spectra of both extracellular and intracellular magnetosomes. XMCD probes the magnetism of a crystal in a site-specific manner, showing that absorption and XMCD ptychography signals provide complementary information. These experiments demonstrate that ptychography, which combines high spatial resolution, high-sensitivity chemical speciation, and site-specific magnetic information, offers a powerful probe for biomineralization studies. Instruments and FacilitiesScanning transmission X-ray microscope (STXM) on beamline 10ID1 of Canadian Light Source at University of Saskatchewan and the Advanced Light Source Beamlines 5.3.2.1 and 11.0.2 at Lawrence Berkeley National Laboratory. | |||
| 11/30/2016 | Crystal Structure of NOV1 Enzyme Reveals Mechanism of Action | Structurally Integrated Biology for the Life Sciences | X-ray Macromolecular Crystallography | Advanced Light Source | Molecular Structure | Stilbenes are compounds produced naturally in a wide variety of plant species and some bacteria, and are also derived from lignin during the kraft pulping process. Enzymes capable of converting stilbenes into useful chemicals or fuels could positively impact the economics of lignocellulosic biomass processing. Collaborators from two U.S. Department of Energy Bioenergy Research Centers report the atomic-level structure of NOV1, a stilbene-cleaving oxygenase (SCO) capable of breaking down the stilbene substrate resveratrol into two smaller compounds. Enzymes such as NOV1 could prove valuable in the biological production of important molecular fragments derived from lignin. | |||
| 11/15/2016 | Mapping the Migration of Genetic Material | National Center for X-Ray Tomography | Soft X-ray Tomography | Advanced Light Source | Cell and Tissue Structure | Researchers used a powerful soft X-ray microscope to capture tomographic images of the genetic material in the nuclei of nerve cells at different stages of maturity. This first quantitative analysis of nuclear organization in intact mammalian cells has generated detailed three-dimensional (3D) visualizations showing an unexpected connectivity in the genetic material called chromatin. The results could aid in understanding how the patterning and reorganization of chromatin relate to the specialization of a stem cell’s function as specific genes are activated or silenced. Understanding the logic of nuclear architecture and how it contributes to stem cell differentiation remains challenging. Until now, imaging the nucleus was possible only indirectly by staining it and hopefully achieving an even distribution. In this work, researchers used soft X-ray microtomography to record a series of images from mouse olfactory nerve cells in three separate stages of development. This unique technique provides a new way of looking at the nucleus without chemically treating the cell, allowing visualization of intact cells in a near-native state at a resolution of about 50 nanometers. The researchers imaged frozen cells at different stages of development from dozens of different angles, then used the images to calculate a 3D reconstruction detailing changing chromatin formations inside the nucleus. The images were collected using soft X-rays within the “water window” (i.e., 284 to 543 electronvolts). The absorption of X-rays by biomolecules is linear with biochemical composition and concentration, generating a unique X-ray linear absorption coefficient measurement for each voxel (i.e., 3D pixel analog). Thus, the researchers were able to visually distinguish between different types of chromatin. Heterochromatin, for example, appears darker than euchromatin in computer-generated tomographic orthoslices (virtual sections) through the nucleus due to its greater biomolecular concentration. The results showed that chromatin compaction increases as the cell matures, and that condensed chromatin moves to the nuclear core during differentiation. Chromatin was thought to exist as a series of disconnected islands, but imaging showed that it compartmentalizes into two distinct regions of “crowding” that form a continuous network throughout the nucleus. Based on a comparison of these results with those of similar cells in which the gene for a heterochromatin binding protein had been inactivated (“knocked out”), the researchers concluded that the protein regulates the reorganization of chromatin in mature neurons. Soft x-ray tomography provides a powerful method to study chromatin and nuclear architecture in vivo. There is no need for chemical fixation or sectioning, preventing a plethora of artifacts introduced by fixatives or by visualization of only thin sections. With the proven success of the imaging technique, the researchers believe it is possible to perform statistical analyses based on large collections of cell nuclei images sorted by different stages of development. Coupled with other types of imaging techniques, researchers hope to isolate individual gene-selection processes in upcoming work. Instruments and FacilitiesSoft X-ray tomography (beamline 2.1) of the Advanced Light Source at the National Center for X-Ray Tomography at Lawrence Berkeley National Laboratory. | |||
| 09/30/2020 | Understanding an Essential Chaperone Complex | Structural Molecular Biology Resource | Cryo-Electron Microscopy, Solution X-ray Scattering, X-ray Macromolecular Crystallography | Advanced Photon Source, National Synchrotron Light Source II, Stanford Synchrotron Radiation Lightsource | Molecular Structure | Ric-8A is a protein involved in the regulation of cell division, which is essential for embryo development. This structure reveals how it acts as a chaperone for Gα in this process. | Scientists reveal a protein binding mechanism connected to embryo development and cancer. The structure of the chaperone protein Ric-8A bound to a G-protein-alpha subunit (Gα) reveals the mechanism for Gα activation through phosphorylation of Ric-8A. | Many modern developments in biology, medicine, and biotechnology have been made possible through our understanding of how biological structures, such as proteins in cells, interact with each other. But to truly reveal their function and the role they play in diseases and medical conditions, scientists need to visualize these structures at the atomic level. This is especially important for more elusive proteins and their functions, such as the Ric-8A protein, a Guanine Nucleotide exchange factor that serves as a chaperone for Gα. Gα are located at membranes and inside cells and are an essential part of several cellular signaling systems. Ric-8A is essential for asymmetric cell division in the process of embryonic development and is also a potential therapeutic target for certain cancers. In a study, scientists revealed the structure and binding mechanism of the Ric-8A protein to the Gαi1 complex. The team used a combination of cryo-electron microscopy and x-ray studies at various light source facilities, including x-ray crystallography at the Frontier Microfocusing Macromolecular Crystallography (FMX) beamline. The FMX beamline is part of the advance life science suite of beamlines at the National Synchrotron Light Source II (NSLS-II), a U.S. Department of Energy (DOE) Office of Science User Facility located at DOE’s Brookhaven National Laboratory. The researchers discovered that the mechanism of Ric-8A differs from the usual mechanism that the G protein uses at its receptors. Ric-8A engages a specific conformation of Gα at multiple interfaces to form a complex that is stabilized by phosphorylation within a Ric-8A segment that connects two Gα binding sites. | |
| 07/31/2020 | Targeting a Critical Molecular Switch in COVID-19 | Center for BioMolecular Structure | Solution X-ray Scattering | National Synchrotron Light Source II | Molecular Structure | This work shows that such inhibitors may possibly be used to fight the COVID-19 pandemic. | Scientists studied the 3-D structure of parts of the COVID-19 virus and found a potential inhibitor for virus replication. They compared the three-stemmed RNA pseudoknots of SARS-CoV and SARS-CoV-2 that function as critical switches in the replication pathways of these viruses and found a small molecule that could inhibit their function. | The COVID-19 pandemic is caused by the severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) and researchers continue to work on treatments and vaccines, many of which can take years to develop. A good short-term strategy may lie in identifying virus-specific targets for small-molecule-based interventions. In this work, a research team investigated how the virus replicates by studying a process called programmed −1 ribosomal frameshift (−1 PRF). In this process, a ribonucleic acid (RNA) pseudoknot acts as a critical switch in the replication programs of the virus and is therefore a promising drug target to inhibit virus replication. The team used small-angle x-ray scattering (SAXS) analyses at the Life Science X-ray Scattering (LiX) beamline at the National Synchrotron Light Source II (NSLS-II) to compare the structure of the three-stemmed pseudoknots of SARS-CoV and SARS-CoV-2. SARS-CoV is a coronavirus strain that caused an epidemic 17 years ago and has been well studied since. NSLS-II is a U.S. Department of Energy Office of Science User facility located at DOE’s Brookhaven National Laboratory. The team showed that the pseudoknots have a similar structure and function. This enabled testing whether a small molecule known to inhibit SARS-CoV frameshifting would be similarly effective against SARS-CoV-2. The results suggest that such frameshift inhibitors may be promising lead compounds to combat the COVID-19 pandemic. | |
| 11/30/2020 | Fine Tuning Drugs to Fight Cancer | Center for BioMolecular Structure, Structural Biology Center | X-ray Macromolecular Crystallography | Advanced Photon Source, National Synchrotron Light Source II | Molecular Structure | Although trametinib is used to treat melanoma, its mechanism of action was not fully understood; this work reveals how the drug binds to its target. | Scientists reveal how the drug trametinib binds itself to its target. Co-crystal structures of the clinical methyl ethyl ketone (MEK) inhibitor, trametinib, demonstrate that the drug binds at the interface of MEK and its scaffold protein, kinase suppressor of Ras (KSR). | A broad range of cancers, including pancreatic, skin, and colorectal cancer, are driven by abnormal activity in the mitogen activation protein kinase network and are linked to frequent mutations in K-RAS and B-RAF. Trametinib is a promising cancer drug that binds to MEK and is currently FDA-approved for the treatment of BRAF V600E/K mutant melanoma; however, how trametinib works and why this drug shows differential activity to other MEK inhibitors has been a mystery. In this work, scientists from the Arvin Dar Laboratory at the Tisch Cancer Institute in the Icahn School of Medicine at Mount Sinai, New York, determined the first ever co-crystal structures of trametinib, revealing an unexpected mode of action whereby the drug spans the interface of an important regulatory complex in the RAS pathway. Using crystallographic, biochemistry, and cell biology analysis, the team found that trametinib simultaneously binds to MEK and its cellular partner KSR. Further, based on these insights, the Dar laboratory designed a novel compound, which they dubbed “Trametiglue,” to overcome a common drug resistance mechanism that limits the efficacy of MEK inhibitors as cancer therapeutics. This work reveals a novel strategy for designing compounds that bind at the interface of protein-protein complexes, which holds great potential for many challenging drug targets. For this study the X-ray diffraction data were collected at the Highly Automated Macromolecular Crystallography (AMX) and Frontier Microfocusing Macromolecular Crystallography (FMX) beamlines at the National Synchrotron Light Source II (NSLS-II). NSLS-II is a U.S. Department of Energy (DOE) Office of Science User Facility located at DOE’s Brookhaven National Laboratory and offers access to state-of-the-art materials characterization and life science tools for researchers from industry and academia. Crystallography data were also collected at the LS-CAT beamline at the Advanced Photon Source at Argonne National Laboratory. | |
| 01/12/2016 | Cellulose Synthesis Complex | Center for Structural Molecular Biology | Small-Angle Neutron Scattering, Solution X-ray Scattering | Spallation Neutron Source/High Flux Isotope Reactor | Molecular Structure | Cellulose is the major structural component of plant cell walls and has great potential as a renewable source of energy. The plant cellulose synthesis complex (CSC), also called a “rosette” because of its hexameric appearance in transmission electron microscope (TEM) images, is a large multi-subunit transmembrane protein complex responsible for synthesis of cellulose chains and their assembly into microfibrils. Despite the importance of cellulose, fundamental properties of the CSC remain unclear. The number of cellulose synthase (CESA) proteins in the CSC and the number of cellulose chains in a microfibril have been debated for years. Vandavasi et al. report a solution structure of the catalytic domain of CESA1 from Arabidopsis thaliana determined by small-angle scattering that provides experimental evidence for the self-assembly of CESA into a stable trimer. This study strongly supports the “hexamer of trimers” model for the rosette CSC that synthesizes an 18-chain cellulose microfibril as its primary product. | |||
| 12/14/2016 | Hollow Pyramid Unlocks Principles of Protein Architecture | Structurally Integrated Biology for the Life Sciences | Solution X-ray Scattering | Advanced Light Source | Molecular Structure | Large biological macromolecules (up to 10,000 atoms) can catalyze chemical reactions in ways that are difficult to replicate inorganically. Their size allows for the complexity needed to perform such functions, but it also makes them more susceptible to misfolding and aggregating—thus, the importance of understanding the fundamental architectural principles that cause large proteins to favor specific conformations. At the nanoscale, where organic chemical groups interact with solvent water molecules, these principles are very different from the ones used to build houses and cars. To investigate these principles, a group of researchers designed a protein that would self-assemble into a hollow pyramid (or tetrahedron). Upon crystallizing the macromolecule, the group found that they had indeed been successful in creating the assembly, but it was unexpectedly warped and collapsed in an asymmetric manner, with some edges bent inward—an asymmetric tetrahedron. To ensure that this was not an artifact of crystallization, they investigated the protein’s behavior in solution using small-angle x-ray scattering (SAXS), which showed that the collapse could be controlled by adjusting the solution’s salt concentration; the structure was disassembled by varying the pH. The flexibility of this macromolecule suggests that it could be useful for the controlled capture and release of smaller compounds. Overall, the researchers expect that, with the tools and techniques developed here, the combination of SAXS with crystallography or electron microscopy could be increasingly useful in analyzing and optimizing designed protein assemblies and understanding their behavior in solution. Instruments and FacilitiesSmall-angle X-ray scattering (SAXS) at Advanced Light Source (ALS) with SIBYLS Beamline 12.3.1 (a joint crystallography and SAXS beamline) at Lawrence Berkeley National Laboratory. | |||
| 12/20/2016 | Detailing the Molecular Roots of Alzheimer’s Disease | Structural Biology Center | X-ray Macromolecular Crystallography | Advanced Photon Source | Molecular Structure | Scientists have captured the three-dimensional (3D) structure of a molecule implicated in Alzheimer’s disease. Knowing the shape of this molecule—and how that shape may be disrupted by genetic mutations—can help in understanding how Alzheimer’s and other neurodegenerative diseases develop and ways to prevent and treat them. Certain mutations that alter the structure of the molecule TREM2 are associated with an increased risk of developing late-onset Alzheimer’s, frontal temporal dementia, Parkinson’s disease, and sporadic amyotrophic lateral sclerosis (ALS), indicating that TREM2 may be involved in cognitive decline. Other TREM2 mutations are linked to Nasu-Hakola disease, a rare inherited condition that causes progressive dementia and death in most patients by age 50. Although its exact contribution is unknown, dysfunctional TREM2 correlates with neurodegeneration, and inflammation appears to be the common thread in neurodegenerative conditions. Structural analysis of TREM2 revealed that the mutations associated with Alzheimer’s alter the surface of the protein, while those linked to Nasu-Hakola influence the “guts” of the protein. The difference in location could explain the severity of Nasu-Hakula, in which signs of dementia begin in young adulthood. The internal mutations totally disrupt the structure of TREM2, resulting in fewer TREM2 molecules. The surface mutations, in contrast, leave TREM2 intact but likely make it harder for the molecule to connect to proteins or send signals as normal TREM2 molecules would. TREM2 lies on the surface of immune cells called microglia, which are thought to be important “housekeeping” cells involved in, for example, maintaining healthy brain biology via a process called phagocytosis that cleans cellular waste, including the amyloid beta that is known to accumulate in Alzheimer’s disease. If the microglia lack TREM2 or the TREM2 is dysfunctional, these cellular housekeepers can’t perform their cleanup tasks. Though the exact function of TREM2 is still unknown, mice without TREM2 have defects in their microglia. With these structural data, scientists can study how TREM2 works, or doesn’t work, in these neurodegenerative diseases, as well as other inflammatory conditions including chronic obstructive pulmonary disease and stroke. The structure of TREM2 could be important for understanding many chronic and degenerative diseases. Instruments and FacilitiesX-ray facility at the Advanced Photon Source (APS) at Argonne National Laboratory. | |||
| 08/15/2016 | How Your Body Transports Zinc to Protect Your Health | Structural Biology Center | X-ray Macromolecular Crystallography | Advanced Photon Source | Molecular Structure | Zinc is essential for wound healing, vision, DNA creation, the senses of taste and smell, and even for sexual health. But despite its importance, scientists have never fully understood the mechanism that moves the mineral through the body—until now. An international collaboration of researchers have, for the first time, created detailed blueprints of the molecular “moving vans” that ferry this important mineral through the blood everywhere it is needed. The finding gives scientists new insights into this important process and a deeper understanding of the critical role it plays in maintaining good health. Zinc is carried through the body by a protein known as serum albumin. Scientists proved the location of the primary site where serum albumin binds with zinc, but the team also found several more secondary binding sites, revealing a more complex interaction than anticipated. While computer models previously had been used to predict how serum albumin picks up zinc, the team used X-ray macromolecular crystallography to obtain data-based images of protein crystals showing how zinc actually bound to serum albumin. The technique allowed them to pinpoint the location of each particular zinc atom, though it was a challenging task. The schematics produced allow scientists to see, for the first time, exactly how serum albumin and zinc come together. Serum albumin also transports many other things, such as hormones and fatty acids. Too much zinc is toxic to the body, so it must make zinc available where it is needed, but prevent harmful, excessive buildup. The study’s findings could help shed light on why certain drugs affect some patients differently than others. Instruments and FacilitiesX-ray macromolecular crystallography; Life Sciences Collaborative Access Team (LS-CAT) 21-ID-F and 21-ID-G X-ray beamlines and Structural Biology Center (SBC)-CAT 19-BM-D X-ray beamline, all at the Advanced Photon Source at Argonne National Laboratory. | |||
| 07/20/2017 | The CRISPR Target-Recognition Mechanism | Structural Molecular Biology Resource | X-ray Macromolecular Crystallography | Advanced Light Source, Stanford Synchrotron Radiation Lightsource | Molecular Structure | Bacterial DNA is characterized by regions of clustered regularly interspaced short palindromic repeats (CRISPRs) and associated Cas proteins (CRISPR-associated endonucleases). The CRISPR-Cas system has revolutionized gene editing by vastly simplifying the insertion of short snippets of new (“donor”) DNA into very specific locations of target DNA. Researchers in this study have discovered how Cas proteins recognize their target locations with such great specificity. They used x-ray crystallography to solve the structures of Cas1 and Cas2—responsible for DNA-snippet capture and integration—as the proteins were bound to synthesized DNA strands designed to mimic different stages of the process. The research also demonstrated how the system works in its native context as part of a bacterial immune system and how Cas proteins act as general-purpose molecular recording devices—tools for encoding information in genomes. Cas1 appears to have evolved from a more “promiscuous” (less selective) type of enzyme that catalyzes the movement of DNA sequences from one position to another (a transposase). At some point, Cas1 acquired an unusual degree of specificity for a particular location in the bacterial genome, the CRISPR array. This specificity is critical to the bacteria, both for acquiring immunity and for avoiding genome damage caused by the insertion of viral fragments at the wrong location. The researchers wanted to learn how Cas1-Cas2 proteins recognize the target sequence to enable comparison with previously studied transposases and integrases (i.e., enzymes that catalyze the integration of donor DNA into target DNA) and to determine whether the proteins can be altered to recognize new sequences for custom applications. To investigate this, they crystallized Cas1-Cas2 in complex with preformed DNA strands that mimicked reaction intermediates and products. X-ray crystallography revealed that the structures showed substantial distortions in the target DNA, but there were surprisingly few sequence-specific contacts with the Cas1-Cas2 complex, and the DNA’s resulting flexibility produced disorder in the crystals. Attempts to model the DNA across the disordered sections showed that the DNA had to be even more distorted. Cryoelectron microscopy experiments, coupled with the crystallography data, confirmed that an accessor protein called the integration host factor (IHF) introduces an additional sharp bend in the DNA, bringing an upstream recognition sequence into contact with Cas1 to increase both the specificity and efficiency of integration. The architecture of the CRISPR integration complex suggests that subtle adjustment of the distance between Cas1 active sites could reprogram the system to recognize different target sites. Changes in its architecture could be exploited, thereby, for genome tagging applications and also may explain the natural divergence of CRISPR arrays in bacteria. Instruments and FacilitiesX-ray macromolecular crystallography; beamline 8.3.1; protein crystallography (PX); and scattering/diffraction at the Advanced Light Source at Lawrence Berkeley National Laboratory; Stanford Synchrotron Radiation Light Source 9-2 beamline. | |||
| 05/03/2018 | Dynamic Regulation of Histone Chaperone Nucleoplasmin | Structural Molecular Biology Resource | Solution X-ray Scattering | Stanford Synchrotron Radiation Lightsource | Molecular Structure | Histones are eukaryotic cell nuclei proteins that package and order DNA into structural units called nucleosomes. Chromatin is the complex of DNA and proteins comprising the genome’s physiological form. As chromatin’s chief protein components, histones act as spools around which DNA winds, playing a role in gene regulation. A chaperone protein assists in the folding and unfolding of macromolecules, such as in the assembly of nucleosomes from folded histones and DNA. Nucleoplasmin (Npm) is a highly conserved embryonic histone chaperone, responsible for the maternal storage and zygotic release of histones H2A and H2B. Npm contains a pentameric N-terminal Core domain and an intrinsically disordered C-terminal Tail domain. Although intrinsically disordered regions are common among histone chaperones, their roles in histone binding and chaperoning have remained unclear. This study, using the Xenopus laevis Npm Tail domain, unveils the architecture of the Npm histone complex and a mechanism of histone chaperone regulation. It demonstrates that intramolecular regulation of the histone chaperone Npm controls histone binding and release—a key process in the earliest stages of embryonic development. Structural analyses enabled model constructions of both the Npm Tail domain and the pentameric complex, revealing that the Tail domain controls the binding of histones through specific, electrostatic interactions. Functional analyses demonstrated that these competitive interactions negatively regulate Npm histone chaperone activity in vitro. Data from these studies establish a potentially generalizable mechanism of histone chaperone regulation via dynamic and specific intramolecular shielding of histone interaction sites. Instruments and FacilitiesBruker 600 nuclear magnetic resonance (NMR) and Inova 600 NMR instruments in the Albert Einstein College of Medicine (AECM) Einstein Structural NMR Resource; and bio–small-angle X-ray scattering (bio-SAXS) beamline 4-2, SLAC National Accelerator Laboratory’s Stanford Synchrotron Radiation Lightsource (SSRL). | |||
| 06/23/2017 | Revealing How Bacterial Organelles Assemble | Structural Molecular Biology Resource | X-ray Macromolecular Crystallography | Stanford Synchrotron Radiation Lightsource | Molecular Structure | Scientists are providing the clearest view yet of an intact bacterial microcompartment, revealing the structure and assembly of the organelle’s protein shell at atomic-level resolution. They studied the “photogenic” organelle shell of an ocean-dwelling slime bacteria Haliangium ochraceum. Providing the first view of the shell of an intact bacterial organelle membrane, this full structural view can help provide important information for beneficial use in fighting pathogens or bioengineering bacterial organelles. The research team said these organelles, or bacterial microcompartments (BMCs), are used by some bacteria to fix carbon dioxide. Thus, understanding how the microcompartment membrane is assembled, as well as how it lets some compounds pass through while impeding others, could contribute to research in enhancing carbon fixation and, more broadly, bioenergy. This class of organelles also helps many types of pathogenic bacteria metabolize compounds that are not available to normal, nonpathogenic microbes, giving the pathogens a competitive advantage. Instruments and FacilitiesMichigan State University–DOE Plant Research Laboratory and the Molecular Biophysics and Integrated Bioimaging Division at Lawrence Berkeley National Laboratory; Stanford Synchrotron Radiation Lightsource. | |||
| 06/26/2017 | Molecular Basis of Alternative Tyrosine Biosynthesis in Plants | Structural Biology Center | X-ray Macromolecular Crystallography | Advanced Photon Source | Molecular Structure | Peanuts have not one, but two ways to make the amino acid tyrosine, an essential plant and human nutrient. Using X-ray crystallography, scientists discovered a mutated form of a key enzyme plants used to make tyrosine. In cherry tomatoes, the basic canonical form of the enzyme dominates; peanuts can switch hit; and some strains of soybeans have lost the canonical form. The discovery demonstrates that primary metabolism can evolve. Researchers have tied this major evolutionary change in plant metabolism to a single mutation in the new enzyme. The more recently evolved of the two pathways for making tyrosine is much less constrained than the earlier one, raising the possibility of using it in commercial production of drugs or nutrients at high yields. Instruments and FacilitiesX-ray macromolecular crystallography; diffraction data collected at beamline 19-ID of the Advanced Photon Source at the Argonne National Laboratory Structural Biology Center. | |||
| 04/12/2017 | Green Alga Genome Provides Blueprint to Advance Clean Energy, Bioproducts | National Center for X-Ray Tomography | Soft X-ray Tomography | Advanced Light Source | Cell and Tissue Structure | Microalgae have potential to help meet energy and food demands without exacerbating environmental problems. The unicellular green alga Chromochloris zofingiensis produces lipids for biofuels and a highly valuable carotenoid nutraceutical, astaxanthin. Thus, advanced understanding of its biology is needed to facilitate commercial development. The assembly of the C. zofingiensis chromosome-level nuclear genome, organelle genomes, and transcriptome from diverse growth conditions was derived from a combination of short- and long-read sequencing in conjunction with optical mapping, revealing a compact genome of ~58 Mbp distributed over 19 chromosomes containing 15,274 predicted protein-coding genes. Found in the genome were 2 genes encoding beta-ketolase (BKT), the key enzyme synthesizing astaxanthin, and both were up-regulated by high light. Isolation and molecular analysis of astaxanthin-deficient mutants showed that BKT1 is required for the production of astaxanthin. Moreover, the transcriptome under high light exposure revealed candidate genes that could be involved in critical yet missing steps of astaxanthin biosynthesis, including ABC transporters, cytochrome P450 enzymes, and an acyltransferase. The high-quality genome and transcriptome provide insight into the green algal lineage and carotenoid production. Microalgae are a promising source of sustainable bioproducts for the increasing demand for food and energy without exacerbating worsening environmental problems. The algae have potential for use as a biofuel feedstock and nutraceutical molecules, including the carotenoid astaxanthin. Analyses of the C. zofingiensis genome and transcriptome and experiments characterizing astaxanthin production advance understanding of algae and carotenoids and enhance the commercial potential of C. zofingiensis. Instruments and FacilitiesSoft X-ray tomography at National Center for X-ray Tomography (NCXT), operated jointly by Berkeley Lab (LBNL) and University of California, San Francisco, at LBNL’s Advanced Light Source. Other techniques: whole-genome optical mapping, high light RNA sequencing, transcriptome sequencing, and long read sequencing. | |||
| 10/06/2016 | Description of Hydration Water in Green Fluorescent Protein Solution | Center for Structural Molecular Biology | Small-Angle Neutron Scattering | Spallation Neutron Source/High Flux Isotope Reactor | Molecular Structure | Many unanswered questions remain about how macromolecules in solution perturb the water molecules in which they are immersed. Scientists used neutron scattering to provide an experimental description of the dynamical perturbation of water molecules surrounding proteins in solution, which play an important role in protein function and stability. Quantifying the magnitude of the perturbation of water around the green fluorescent protein (GFP), they found a systematic length-scale dependence of the dynamical retardation factor compared to the perturbation of bulk water. The findings deepen the understanding of water perturbation, which is of practical interest to researchers in food science, personal care, pharmaceutics, and protein dynamics. Instruments and FacilitiesNeutron scattering at Center for Structural Molecular Biology and Spallation Neutron Source at Oak Ridge National Laboratory. | |||
| 09/29/2017 | Precise Control of Neutron Contrast in Surfactant Micelles Provides Platform for Membrane Structure Studies | Center for Structural Molecular Biology | Small-Angle Neutron Scattering | Spallation Neutron Source/High Flux Isotope Reactor | Molecular Structure | Scientists in this study have successfully demonstrated the ability to manipulate the neutron contrast of detergent micelles by incorporating a similar detergent with deuterium-labelled alkyl chains. The presence of excess detergent micelle scattering often has a detrimental influence on scattering data obtained for membrane protein–detergent complexes. Isolation of the scattering signal from the protein of interest can be accomplished by eliminating all scattering from the detergent. This approach enabled determination of the overall structure and oligomeric state of a small membrane protein enzyme. Instruments and FacilitiesSmall angle neutron scattering (SANS): Bio-SANS beamline (CG3) of the High-Flux Isotope Reactor at Oak Ridge National Laboratory (ORNL). Recorded scattering data using MantidPlot software. Neutron contrast studies: ModULes for the Analysis of Contrast (MULCh) Variation Data at the University of Sydney. | |||
| 12/22/2016 | Neutrons Identify Oxygen Activation in LPMO enzyme | Center for Structural Molecular Biology | Neutron Macromolecular Crystallography, X-ray Macromolecular Crystallography | Advanced Photon Source, Spallation Neutron Source/High Flux Isotope Reactor | Molecular Structure | Fungal lytic polysaccharide monooxygenases (LPMOs) are known to enhance the efficiency of cellulose-hydrolyzing enzymes through oxidative cleavage of the glycosidic bonds. For this study, PMO-2 from Neurospora crassa was heterologously expressed from Pichia pastoris, purified, and crystallized for high-resolution X-ray crystal structures that revealed “prebound” molecular oxygen in the resting state and a dioxo species in complex with the catalytic copper (Cu2+) ion, which is the first structural description of molecular oxygen (O2) activation by a LPMO. In addition, neutron diffraction studies and density functional theory calculations have identified a role for a conserved histidine in promoting oxygen activation. Extension of these studies to the enzyme-substrate complex could provide a complete picture of the enzymatic mechanism for the potential benefit of applications such as bioethanol production. Instruments and FacilitiesNeutron crystallography. Joint X-ray/neutron refinement at Center for Structural Molecular Biology at Oak Ridge National Laboratory (ORNL). Diffraction data were collected at SER-CAT 22-ID at the Advanced Photon Source at Argonne National Laboratory and at CG-4D IMAGINE (NSF MRI 09229719) at the High Flux Isotope Reactor at ORNL. | |||
| 05/23/2017 | Using Neutrons to Resolve Plasma Membrane Organization in Live Bacteria | Center for Structural Molecular Biology | Small-Angle Neutron Scattering | Spallation Neutron Source/High Flux Isotope Reactor | Molecular Structure | A new strategy devised by scientists at Oak Ridge National Laboratory used nondestructive small-angle neutron scattering (SANS) to reveal, for the first time, nanoscale membrane structures in living cells. Neutron scattering spectra confirmed that the plasma membrane of the bacterium Bacillus subtilis is lamellar with an average hydrophobic thickness of 24 Ångstroms. The data also revealed that the membrane contains lipid features of approximately 40 nm or less in size, consistent with hypothesized “lipid rafts” in biological systems. The observation of lipid segregation in the plasma membrane of a bacterium is consistent with the notion of nanoscopic lipid assemblies, often described as lipid rafts in mammalian systems, implying that lipid domains are integral features of all biological membranes. Among their functions, lipid rafts are thought to play a vital role in cell signaling and facilitate movement of essential biomolecules in and out of the cell. In addition to these scientific findings, the methods developed provide a new experimental platform for pursuing additional areas of inquiry (e.g., systematic in vivo investigations of cell membrane structure and response to diverse environmental stimuli). This new approach also may prove valuable, for example, in biomass feedstock and biofuel production, where bacterial cell membranes play important roles, and in biomedicine, where bacterial membrane domains affect antibiotic resistance. Furthermore, the strategy for “visualizing” the membrane can be used with other physical characterization techniques to examine additional cell structures such as the cell wall. Instruments and FacilitiesSmall-angle neutron scattering (SANS) at the High Flux Isotope Reactor (HFIR) at Oak Ridge National Laboratory (ORNL). EQ-SANS at Spallation Neutron Source at ORNL. Oak Ridge Leadership Computing Facility at ORNL. | |||
| 08/03/2017 | Chemical Strategy for Conversion of CO2 to Biomass | Structural Biology Center, Structural Molecular Biology Resource | X-ray Macromolecular Crystallography | Advanced Photon Source, Stanford Synchrotron Radiation Lightsource | Molecular Structure | A research team has uncovered a new mechanism for enzyme-mediated carbon dioxide (CO2) capture and conversion. They have revealed new and unique elements of carboxylation chemistry (CC) that could be used in catalytic strategies for converting CO2 into chemical feedstock or biomass. The crystal structure of acetone carboxylase (AC; enzyme involved in biodegradation by bacteria) was solved using X-ray crystallographic data. The inactive, unbound (apo) structure shows a substrate channel blocked off from the manganese (Mn) active site where CO2 conversion takes place. An adenosine monophosphate (AMP)–bound structure contains large conformational changes that open the channel to the active site. Highly reactive intermediates in the channel are protected from outside solvent as they are transported to the Mn active site for CO2 conversion. Stepwise mechanisms of carboxylation reactions differ in essential ways with respect to co-substrate, co-factor, and metal requirements. Knowledge of these mechanisms provides the basis for an increased fundamental understanding of CC, contributing to future strategies for CO2 capture and conversion to biomass. These in turn may mitigate the effects of increasing concentrations of CO2 on the global climate. Instruments and FacilitiesX-ray crystallographic data from the Stanford Synchrotron Radiation Lightsource (SSRL) at SLAC National Accelerator Laboratory (SLAC). Structural data from X-ray macromolecular crystallography were measured on SSRL beamline 12-2; Advanced Photon Source (APS) at Argonne National Laboratory (ANL); and the Diamond Light Source, Oxfordshire, United Kingdom. | |||
| 11/19/2020 | Enhancing Enzyme Catalysis Through Directed Evolution | Structural Molecular Biology Resource | X-ray Macromolecular Crystallography | Stanford Synchrotron Radiation Lightsource | Molecular Structure | Computational enzyme design has aided the development of catalysts for chemical reactions ranging from ester hydrolysis to cycloadditions. Although the starting activities for these enzymes are usually low, they can be increased to levels approaching those of natural enzymes through laboratory evolution. This process mimics the natural selection of enzymes in biology, with the advantage that individual intermediates along the evolutionary pathway can be characterized to deduce how function was enhanced. For the computationally designed Kemp eliminase HG3 enzyme, researchers wanted to better understand the molecular origins of a rate enhancement achieved by directed evolution. The Kemp elimination, a well-studied model based on a basic xylanase scaffold for proton transfer from carbon, has served as a benchmark for de novo enzyme design. The research team—which included collaborators from the Swiss Federal Institute of Technology, Brandeis University, University of Massachusetts, and Stanford Synchrotron Radiation Lightsource (SSRL)—obtained two evolutionary intermediates from HG3: HG3.7 after seven rounds of mutagenesis and screening and HG3.17 after 17 rounds. The team analyzed these intermediates using nuclear magnetic resonance and x-ray crystallography. The HG3.17 intermediate showed a 200-fold increase in reaction rate compared to HG3. Researchers determined crystal structures of HG3, HG3.7, and HG3.17 using diffraction data collected at SSRL beamlines 14-1 and 7-1 at multiple temperatures and within novel graphene crystal mounts. The team found that HG3 and HG3.7 both had dual conformers near the active site, one of which can destabilize the transition state. However, in HG3.17, only one confomer existed, which stablizes the transition state. This study shows that explicit screening of productive conformations that confer better enzymatic activity can be used to improve design protocols and guide development of future mutagenesis and screening strategies. | |||
| 08/14/2020 | Phenoxy Radical-Radical Coupling in Plants | Structural Molecular Biology Resource | X-ray Macromolecular Crystallography | Stanford Synchrotron Radiation Lightsource | Molecular Structure | To better understand DP biochemical activities and how they give rise to distinct classes of plant phenolic compounds, research teams examined DP structures using the capabilities of the Structural Molecular Biology program at the Stanford Synchrotron Radiation Lightsource (SSRL). Results will help inform predictions and efforts to identify the metabolic pathways and precise biochemical roles that DPs play in the coupling reactions that lead to important plant compounds. | The evolution of phenolic radical-radical coupling in plants was pivotal in the transition of plants from early aquatic forms, such as algae and cyanobacteria, to the huge diversity of terrestrial plants today. The reactions form carbon-carbon and carbon-oxygen bonds via radical formation in phenolate compounds, and the molecules ultimately produced are responsible for defense, structural integrity, and reinforcement of plants ranging from grasses and ferns to the largest trees. These reactions generate compounds such as lignans, lignins, pterocarpans, and gossypol, which together account for 30% to 50% of vascular plant mass and represent a massive sink for sequestered carbon. Research to understand the formation of these compounds has found that the first step in lignan biosynthesis is the stereospecific coupling of two molecules of coniferyl alcohol. Studies revealed two isomers of the lignan product pinoresinol in plants, but how these reactions were catalyzed was not fully understood until researchers from Washington State University identified dirigent proteins (DPs) in the late 1990s. DPs evidently emerged during plants’ transition from water to land, with phylogenetic analyses revealing numerous DP subfamilies throughout the plant kingdom. The biochemical functions of most of these DPs (>95%) still are unknown. | In 2015, researchers used data collected at SSRL beamline 12-2 to solve the structure of the DP that forms (+)-pinoresinol, a widely distributed plant lignan (Kim et al. 2015). The structure was solved as a dimer of two β-barrel molecules in the asymmetric unit, but there is a very tight trimer generated by a threefold axis. Despite the oligomeric unit most likely being a trimer, each monomer acts independently, and each has an isolated active site at one end of the β-barrel. Recently, Meng et al. (2020) used data collected at beamline 9-2 to determine the crystal structures of two additional DPs that catalyze the formation of medicarpin, a pterocarpan precursor (see figure). Docking simulations and analysis of the substrate binding sites show how the diastereomeric chiral isoflavonoid precursors might bind and how the radical formed during the subsequent coupling reaction would be stabilized. Moreover, Washington State University, SSRL, Pacific Northwest National Laboratory, and the U.S. Department of Energy’s Joint Genome Institute recently established a joint research program to generate a large number of DPs of varying function. Each DP will be characterized enzymatically and structurally to gain a better understanding of the range and variability of the coupling reactions catalyzed by these important enzymes. | |
| 06/22/2017 | Archaeal Abundance on Human Skin Changes with Age | Berkeley Synchrotron Infrared Structural Biology Imaging Program | Synchrotron Infrared Hyperspectral Imaging | Advanced Light Source | Chemical and Elemental Information | A 2017 study discovers archaea among the billions of bacteria comprising the human skin microbiome, and that their abundance varies with a person’s age. To characterize the microbes that live on human skin, researchers from Austria collaborated with Hoi-Ying Holman and other beamline scientists at the Berkeley Synchrotron Infrared Structural Biology Imaging Program, a BER-supported infrared beamline at the Advanced Light Source at Lawrence Berkeley National Laboratory. The research was a joint project between NASA and the European Space Agency. Based on the chemical specificity of infrared spectroscopy, the infrared beamline rapidly and precisely characterized samples from the skin of human volunteers aged 1 to 75 years and determined the levels and types of microbes present. The analysis was then linked back to genomic data collected by the Austrian team, which revealed the presence of archaea. Before the study, the existence of archaea on human skin was unknown. The genetic and chemical analyses of samples collected from showed that archaea were most abundant in subjects younger than 12 and older than 60, and in people with dry skin. Archaeal abundance was not associated with sex. The detected archaea are thought to be involved in nitrogen turnover and can lower skin pH, which helps suppress pathogens and prevent infection. Instruments and FacilitiesFluorescence in situ hybridization (FISH); quantitative polymerase chain reaction (PCR); next-generation sequencing; Fourier Transform infrared (FTIR) focal plan array (FPA) hyperspectral imaging. Facilities: Advanced Light Source at the Berkeley Synchrotron Infrared Structural Biology Imaging Project; BioTechMed-Graz, the Bavaria California Technology Center. | |||
| 08/11/2017 | Hacking the Bacterial Social Network | Structural Biology Center | X-ray Macromolecular Crystallography | Advanced Photon Source | Molecular Structure | Scientists have determined the molecular structures of a highly specialized set of proteins that a strain of Escherichia coli bacteria use to communicate and defend their turf. The research shows that bacteria appear to have a social network that lets them attract and repel one another, potentially leading to new biomedical strategies for overcoming pathogenic bacteria that cause infectious diseases such as pneumonia and food-borne illnesses. The work builds on the 2005 discovery that bacteria produce toxic proteins which they can transfer to their neighbors through direct contact to either kill or control them. This strategy may help them gain better access to nutrients in densely populated microbial communities through a process called contact-dependent growth inhibition (CDI). Learning how bacteria interact and communicate is helping to resolve the different activities of the toxins, which may affect different bacteria differently. CDI system toxins are found in soil and gut bacteria, as well as in human pathogens like Pseudomonas aeruginosa which is involved in lung disease. The research team used high-bright X-rays to characterize biological proteins and inspect chemical processes at the nanoscale level, obtaining the molecular structures of proteins that belong to a three-part system of the NC101 strain of E. coli. The system consists of the CDI toxin, its immunity protein, and its elongation factor (EF). The latter, known as EF-Tu, is a protein that plays a key role in protein synthesis. Knowing the protein structures of all three parts helps scientists understand their function. Discovery of the immunity protein has led scientists to suspect that the system’s purpose includes competition and signaling (intracommunication), as well as killing and controlling other bacteria. Only a few molecules of the toxin reach the neighboring cell, so perhaps the toxin is not meant to kill, but rather to control and communicate. The toxin can act on transfer ribonucleic acid (tRNA) only under highly specific circumstances and it is the first case seen where EF is needed for the toxin to function. Instruments and FacilitiesAdvanced Photon Source 19-ID beamline and Advanced Protein Characterization Facility at Argonne National Laboratory. | |||
| 06/16/2017 | Engineering Better Plant Cultivars for Iron Uptake by Modifying the Iron Deficiency Response in <i>Arabidopsis thaliana</i> | Structural Molecular Biology Resource | X-ray Fluorescence Imaging | Advanced Photon Source, National Synchrotron Light Source II, Stanford Synchrotron Radiation Lightsource | Chemical and Elemental Information | Many populations in developing countries rely on plants for dietary iron (Fe) which is essential for plant growth, crop yields, and human health. However, due to its low or limited solubility, Fe is sparingly available in neutral or basic soils and thus not readily accessible in the rhizosphere. This low solubility leads to restricted Fe content in many plants and is a major factor contributing to the widespread prevalence of Fe deficiency anemia in people with plant-based diets. Increasing plant Fe acquisition and storage may have profound impacts on plant and human nutrition and can be achieved by manipulating genes and related mechanisms governing Fe homeostasis in plants. However, understanding the balance between positive and negative regulation of the Fe deficiency response is essential for efforts to engineer plants having a sufficient but not toxic level of Fe. Although plants often are challenged with Fe deficiency, no environment remains constant, making Fe availability in the rhizosphere dependent on many factors. When sufficient Fe is available, plants must effectively suppress their Fe-deficiency response to avoid excessive uptake. A research team has identified a novel Fe-binding domain allele called BTS in a mutagenesis screen for altered Fe accumulation (bts-3). Data showed that bts-3 is more tolerant than wild type to Fe-deficient conditions and that bts-3 is sensitive to Fe-sufficient conditions and accumulates excessive Fe. A triple mutant with loss of both BTS paralogs and a partial loss of BTS expression exhibits even greater tolerance to Fe-deficient conditions and increased Fe accumulation without any resulting Fe toxicity effects, with the mutations also changing their uptake of important minerals such as zinc (Zn) and manganese (Mn). Genetic knockdowns and modifications of the proteins have been implicated in regulating plant uptake of Fe. This work will lead to greater understanding of plant Fe homeostasis to inform efforts for improved crops. Identifying natural variants of these genes in crop species may lead to traditional breeding efforts to generate higher-Fe cultivars. Instruments and FacilitiesPerkinElmer LAS Ltd, Seer Green, United Kingdom, and Elemental Scientific Inc., Omaha, Neb.; synchrotron X-ray fluorescence (SXRF) at National Synchrotron Light Source (NSLS) at Brookhaven National Laboratory; Stanford Synchrotron Lightsource (SSLS) at SLAC National Accelerator Laboratory (SLAC); real-time quantitative PCR (Step One Plus Real Time PCR System using Applied Biosystems Version 2.2.3; Advanced Photon Source (APS) at Argonne National Laboratory (ANL); and Australian Synchrotron, Victoria. Two-dimensional SXRF analysis was performed at various X-ray microprobe beamlines: microarray analysis and X-ray fluorescence imaging (XFI) on SSLS beamline 2-3 at SLAC; X26A and X27A of NSLS; XFM beamline of the Australian Synchrotron; APS beamline 2-ID-D. Microarray analysis performed at Geisel School of Medicine in the Genomics Shared Resource at Dartmouth College. Elemental concentration analysis (inductively coupled plasma-mass spectrometry (ICP-MS, PerkinElmer NexION 300D equipped with Elemental Scientific Inc. autosampler and Apex HF sample introduction system at PerkinElmer LAS Ltd, Seer Green, U.K., and Elemental Scientific Inc., Omaha, Neb., respectively, in the standard mode. | |||
| 05/09/2017 | Probing S-layer Protein Structural Dynamics with SAXS | Structural Molecular Biology Resource | Solution X-ray Scattering | Stanford Synchrotron Radiation Lightsource | Molecular Structure | All archaea and many bacteria possess a protein shell referred to as a surface layer (S-layer), which usually consists of a single protein self-assembled into a two-dimensional (2D) crystal lattice. Studies have revealed the structural dynamics of this S-layer protein from the model bacterium Caulobacter crescentus, called RsaA. 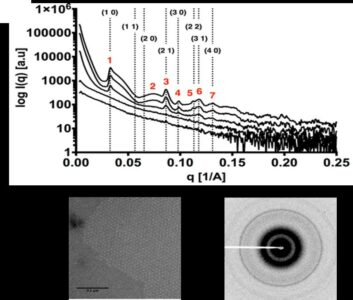 Small angle X-ray scattering and diffraction (SAXS/D) data of five solutions with different concentrations of the Caulobacter crescentus S-layer protein, RsaA, in the presence of calcium (Ca). Scattering profiles indicate concentration-dependent crystallization. Automatic indexing of the numbered peaks yielded a hexagonal crystal lattice consistent with predictions and denoted by Miller indices. (Top) The diffraction pattern obtained for the highest concentration used (8 mg/ml) shows powder rings. (Bottom) Transmission electron microscopy of the 8 mg/ml RsaA in the presence of 10 millimole per Liter (mm/L) of calcium chloride (CaCl2) (scale bar 200 nm). [Reprinted from Herrmann, J., et al. (2017). DOI:10.1016/j.bpj.2017.04.003. Copyright 2017, with permission from Elsevier.] Enabling differentiation of the discrete states, these results rationalize physiological data implicating RsaA as a player in environmental adaptation of C. crescentus. The findings also provide a biochemical and physiological basis for RsaA’s calcium (Ca)-binding behavior, which extends far beyond Ca’s usual role in S-layer biology of aiding biogenesis or oligomerization, and demonstrate a connection to cellular fitness. Further characterization using slow and fast time-resolved SAXS/D methods is ongoing. Instruments and FacilitiesStanford ChEM-H Macromolecular Structure Knowledge Center, Stanford Department of Structural Biology Electron Microscopy Center, Stanford Synchrotron Radiation Lightsource (SSRL) at SLAC National Accelerator Laboratory (SLAC). Beamlines or instruments used: transmission electron microscopy (TEM) and small angle X-ray scattering and diffraction (SAXS/D) at SSRL beamline 4-2 at SLAC. | |||
| 09/08/2017 | The Origins of Photosynthesis in a Sun-Loving Bacteria | Structural Biology Center | X-ray Macromolecular Crystallography | Advanced Light Source, Advanced Photon Source | Molecular Structure | The simplest known bacterium able to drive photosynthesis is found in muddy soils near hot springs. Heliobacterium modesticaldum is a sun-loving, soil-dwelling, thermophilic bacterium that photosynthesizes near-infrared light, unlike plants, which use different parts of visible light. The photosynthesis reaction centers (RCs) of H. modesticaldum are thought to resemble the earliest common ancestor of all photosynthesis complexes, which evolved around three billion years ago. Thus, a clear, detailed picture of the H. modesticaldum RC would provide valuable insight into the early evolution of photosynthesis. However, the process to successfully purify an RC protein and grow the crystals needed for X-ray crystallography can be lengthy and difficult. Researchers successfully capped 7 years of work by obtaining a high-resolution (2.2 Å) structure of the membrane protein at the heart of the photosynthetic RC of H. modesticaldum. The structure revealed details such as the interactions between light-harvesting “antenna” molecules and the electron-transfer chain, where the initial steps required to convert photon energy into chemical energy occurs. With this data, the research team was able to make comparisons to other types of RCs, gaining new perspectives on the early evolution of photosynthesis and how nature optimized light-driven energy collection. This work will help unlock the secrets of photosynthesis, possibly leading to the development of cleaner, solar-based renewable energy. Instruments and FacilitiesX-ray macromolecular crystallograpy at the Biodesign Center for Applied Structural Discovery at Arizona State University; Beamline 8.2.1 at the Advanced Light Source at the Berkeley Center for Structural Biology at Lawrence Berkeley National Laboratory; the Structural Biology Center–CAT beamline of the Advanced Photon Source at Argonne National Laboratory. | |||
| 08/14/2017 | X-ray Footprinting Solves Mystery of Metal-Breathing Protein | Center for BioMolecular Structure | X-ray Footprinting | Advanced Light Source | Molecular Structure | Scientists have discovered the details of an unconventional coupling between a protein from the metal-reducing bacterium Shewanella oneidensis and a mineral that allows the bacterium to breathe when oxygen is unavailable. The work could help answer a long-standing question in microbiology: how do bacterial proteins interact directly with minerals to transfer electrons and allow the microbe to live? The researchers used an X-ray-based technique known as “footprinting” to pinpoint the chemical connections between the bacterial protein and nanoparticles composed of iron and oxygen. Footprinting reveals interactions of proteins in a near-native environment. The technique identified a surprisingly small and weak binding site that had already been mapped, but it also revealed how the site binds to metal-containing minerals—a feat that conventional techniques had been unable to accomplish. The study provides insight into ways to design proteins with better electronic connections that can be used to build more sensitive bioelectronic sensors. It could also lead to new innovations in linking proteins to other materials for bio-based electronic devices, such as sensors that can diagnose disease or detect contaminants. And it could help researchers understand and control the chemical reactions sparked by these protein-material interactions and, eventually, how organisms remodel their environment and make biominerals. Instruments and FacilitiesX-ray footprinting was performed using X-ray beamline 5.3.1 at Lawrence Berkeley National Laboratory’s Advanced Light Source (ALS), followed by mass spectrometry at the Biological Nanostructures Facility at the Molecular Foundry. | |||
| 12/05/2016 | Designing Cyclic Oligomers: Greater than the Sum of Their Parts | Structurally Integrated Biology for the Life Sciences | Solution X-ray Scattering, X-ray Macromolecular Crystallography | Advanced Light Source, Advanced Photon Source | Molecular Structure | Cyclic proteins that assemble from multiple identical subunits (homo-oligomers) play key roles in many biological processes, including enzymatic catalysis and function and cell signaling. Researchers in the Molecular Biophysics and Integrated Bioimaging (MBIB) Division worked with University of Washington’s David Baker, who led a team to design in silico and crystallize self-assembling cyclic homo-oligomer proteins. A strategy was developed that designs interfaces onto idealized proteins aimed to direct their assembly into multimeric complexes. Researchers used structural characterization, both X-ray crystallography and small angle X-ray scattering (SAXS), to show that many of the designs adopted the target oligomerization state and predicted structure. Not only does the work demonstrate that scientists have a basic understanding of what determines oligomerization, but that they could design proteins with tunable shape, size, and symmetry for a variety of biological applications. Instruments and FacilitiesCrystallography Collective program; beamline 5.0.2; small angle X-ray scattering (SAXS) on the SIBYLS BL12.3.1 beamline. Advanced Photon Source synchrotron beamline 24-ID-C at Berkeley Center for Structural Biology and Advanced Light Source at Lawrence Berkeley National Laboratory (LBNL). | |||
| 09/09/2017 | Measuring and Modeling Poplar Root Water Extraction After Drought Using Neutron Imaging | Center for Structural Molecular Biology | Neutron Imaging | Spallation Neutron Source/High Flux Isotope Reactor | Cell and Tissue Structure | Neutron radiography was used to measure soil water movement and water uptake by individual roots in situ. Root water uptake was linked to root traits; smaller-diameter roots had greater water uptake per unit surface area than larger-diameter roots. Model analysis based on root-free soil hydraulic properties indicated unreasonably large water fluxes among the vertical soil layers during the first 16 hours after wetting. These results suggest problems with common soil hydraulic or root surface area modeling approaches, indicating the need to further investigate and understand the impacts of roots on soil hydraulic properties. This work highlights the team’s ability to link root water uptake to characteristic root traits, thus enabling performance assessment of common water uptake models. Instruments and Facilities UsedSequential neutron radiography using CG1-D beam line using cold neutrons, at High Flux Isotope Reactor (HFIR) at Oak Ridge National Laboratory. Neutron attenuation by plant samples was detected with a 25-µm lithium fluoride/zinc sulfide (LiF/ ZnS) scintillator linked to a charge coupled detector (CCD) camera system (iKon – L 936, Andor Technology plc., Belfast, U.K.). Roots scanned and dimensions measured using WinRhizo software (Regent Instruments Inc., Quebec, Canada). | |||
| 06/30/2017 | Finding New Clues to a Common Respiratory Virus | Structural Biology Center | X-ray Macromolecular Crystallography | Advanced Photon Source | Molecular Structure | Each year in the United States, more than 57,000 children younger than 5 years old are hospitalized due to respiratory syncytial virus (RSV) infection, and about 14,000 adults older than 65 die from it. By age 2, most children have been infected with RSV, experiencing usually only mild cold symptoms. People with weakened immune systems, however, such as infants and the elderly, can face serious complications, including pneumonia and, in some cases, death. Now, scientists studying the virus using bright X-rays have found clues to how RSV causes disease. They mapped the molecular structure of an RSV protein that interferes with the body’s ability to fight off the virus. Knowing the structure of this protein will help them understand how the virus impedes immune response, potentially leading to a vaccine or treatment for this common infection. With no approved vaccine and limited treatment for RSV, doctors prescribe the antiviral drug ribavirin only in the most severe cases because it is expensive and not very effective. Thus, most people with RSV receive only supportive care to make them more comfortable while their bodies fight off the virus. People with weakened immune systems face a tough fight with RSV. The researchers say that solving the structure of this elusive protein will enable them to see what the protein looks like and help them define what it does and how it does it. Ultimately, this capability and finding could lead to new targets for vaccine or drug development. Scientists have long known that a nonstructural RSV protein (known as NS1) is key to the virus’s ability to evade the body’s immune response. However, its structure was unknown, so scientists were unable to determine exactly how the enigmatic NS1 interfered with the immune system. Using X-ray crystallography, the scientists determined the three-dimensional (3D) structure of NS1, and, in a detailed analysis of the structure, they identified a piece of the protein known as the alpha 3 helix, possibly critical for suppressing the immune response. The researchers created different versions of this NS1 protein, with some having an intact and some with a mutated alpha 3 helix region. They tested the functional impact of helix 3 and created a set of viruses containing either the original or the mutant NS1 genes, measuring the effect on the immune response when they infected cells with these viruses. They found that viruses with the mutated helix region did not suppress the immune response, while the ones with the intact helix region did, globally modulating the immune response. The findings also showed that the protein’s alpha 3 helix region was necessary for the virus to dial down the body’s immune response, giving the virus a better chance of surviving and multiplying—in other words, causing disease. Thus, a vaccine or treatment that targets the alpha 3 helix may supply what is needed to prevent immune suppression. | |||
| 11/02/2017 | Arsenic Accumulation in Benthic Periphytic Biofilms | Structural Molecular Biology Resource | X-ray Fluorescence Imaging | Stanford Synchrotron Radiation Lightsource | Chemical and Elemental Information | Arsenic (As) is a highly toxic metalloid that occurs naturally in the Earth’s crust and can be mobilized as a result of anthropogenic activities. Arsenic can exist in two different oxidation states, As(III) and As(V). These different chemical forms have varying mobilities and bioavailabilities. Understanding the biogeochemical controls on As speciation is therefore necessary to predict how As will accumulate in aquatic food webs. Benthic periphytic biofilms form on streambeds and have been shown to accumulate high concentrations of As. Furthermore, periphytic biofilms are at the base of the food chain in aquatic environments and thus could transfer accumulated As to organisms at higher trophic levels. Lopez et al. sought to determine how inorganic As(III) and As(V) were taken up by periphytic biofilms, how these biofilms modified As speciation, and whether these biofilms were a source of As to higher trophic levels. In this study, Lopez et al. examined the uptake of As(III) versus As(V) by periphytic biofilms. They also fed these biofilms to larval mayflies. Both oxidation states of As were taken up by the biofilms, leading to ~6,000 times higher As concentrations relative to the initial solutions. However, the concentrations present in the larval mayflies were diluted relative to the biofilms (for both oxidation states), showing that As was not biomagnified in this food chain and suggesting that As was sequestered in the biofilm in a form that was not bioavailable. The authors used X-ray fluorescence imaging performed on beamline 2-3 at the Stanford Synchrotron Radiation Lightsource (SSRL) to examine the speciation of As in periphytic biofilms under their different experimental conditions. This showed that (1) As was associated with iron regardless of its initial oxidation state and (2) As(III) was oxidized to As(V) when taken up by the periphyton biofilm. The authors t concluded that the periphyton biofilm sequestered As by oxidizing As(III) to As(V), possibly via redox active iron or manganese oxides present within the biofilm. The As(V) then adsorbed strongly to iron oxides. In summary, Lopez et al. have demonstrated the ability of benthic periphytic biofilms to accumulate As(V) and As(III) efficiently, without transferring As to higher trophic levels. This likely was because As was sequestered in a strongly adsorbed As(V) complex on iron oxide surfaces. | |||
| 10/01/2016 | Crystal Clear Imaging Infrared Brings to Light Nanoscale Molecular Arrangement | Berkeley Synchrotron Infrared Structural Biology Imaging Program | Synchrotron Infrared Hyperspectral Imaging | Advanced Light Source | Chemical and Elemental Information | A team of researchers from Lawrence Berkeley National Laboratory (LBNL) and the University of Colorado–Boulder have developed a new way to reveal crystal features in functional materials for understanding not only their composition but also their molecular arrangement and microscopic imperfections. Using LBNL’s Advanced Light Source (ALS), they combined the power of infrared light from the ALS and infrared light from a laser with an atomic force microscope to reveal key nanoscale details about the nature of an organic semiconductor, revealing its crystal shapes, orientations, and defects that affect its performance. | |||
| 07/10/2017 | Novel Orange Carotenoid Proteins Shed Light on Evolution of Cyanobacteria Photoprotection | Structurally Integrated Biology for the Life Sciences | X-ray Macromolecular Crystallography | Advanced Light Source | Molecular Structure | Research has identified and characterized a new, functionally distinct member of the orange carotenoid protein (OCP) family. The OCP complex enables chromatically acclimating blue-green algae to avoid cellular damage and growth inhibition in conditions of high light or nutrient stress. In a recent bioinformatic analysis of all available cyanobacterial genomes, scientists found that many of these ecophysiologically diverse organisms encode more than one copy of the full-length OCP. The study’s focus was the filamentous blue-green algae Tolypothrix, which encodes two OCPs. One copy was determined to be functionally equivalent to the well-characterized OCP of Synechocystis cyanobacteria, dubbed OCP1. But the second, OCP2, was distinct in several key aspects. The researchers hypothesize that OCP2 and another bioinformatically identified protein, OCPx, reflect intermediate stages in the evolution of photoprotection in cyanobacteria. Instruments and FacilitiesX-ray macromolecular crystallography and diffraction was conducted at the Advanced Light Source at Lawrence Berkeley National Laboratory. | |||
| 01/11/2017 | Study Pinpoints Most Reactive Areas of Nanoscale Particles | Berkeley Synchrotron Infrared Structural Biology Imaging Program | Synchrotron Infrared Hyperspectral Imaging | Advanced Light Source | Chemical and Elemental Information | Key hot spots for chemical reactivity in nanosized platinum and gold particles have been found in the defects and jagged surfaces at their edges. Understanding and customizing the structural properties of these catalysts are important in the production of many industrial products, such as fertilizers, fuel, and plastics, to make them more efficient while remaining unchanged in the process. A team of researchers working at the U.S. Department of Energy’s Lawrence Berkeley National Laboratory (LBNL) and the Hebrew University of Jerusalem in Israel confirmed how the atomic structure of nanoparticles affects their function with a unique infrared probe at LBNL’s Advanced Light Source. | |||
| 01/19/2017 | A Regional Model for Uranium Redox and Mobility | Structural Molecular Biology Resource | X-ray Absorption and Emission Spectroscopy | Stanford Synchrotron Radiation Lightsource | Chemical and Elemental Information | Uranium (U) contamination stubbornly persists as a challenging and costly water quality concern at former uranium ore processing sites across the Upper Colorado River Basin (UCRB). Plumes at these sites are not self-attenuating via natural flushing by groundwater as originally expected. Recent studies at the Rifle, Colo., legacy site suggest that organic-enriched anoxic sediments create conditions that promote reduction of U(VI) to relatively immobile U(IV), causing it to accumulate locally under persistently saturated and anoxic conditions. However, incursion of oxidants into reduced sediments could transform contaminants, allowing these sediments to act as secondary sources of uranium. Oxidant incursions take place during periods of changing water tables, which occur in UCRB throughout the year. If these sediments were regionally common in the UCRB and exposed to varying reduction-oxidation (redox) conditions, then they could contribute to maintaining the longevity of regional uranium plumes. To investigate these issues, researchers examined the occurrence and distribution of reduced and oxidized iron (Fe), sulfur (S), and U species in sediment cores spanning dry and oxic to wet and reduced conditions at three different UCRB sites. Detailed molecular characterization involved chemical extractions, X-ray absorption spectroscopy (XAS), Mössbauer spectroscopy, and X-ray microspectroscopy. This work demonstrates that anoxic organic-enriched sediments occur at all sites, strongly accumulate sulfides and U, and are exposed to strong seasonal redox cycles. Uranium was found to be present as U(IV) complexed to sediment-associated organic carbon and possibly to mineral surfaces. This finding is significant because complexed U(IV) is relatively susceptible to oxidative mobilization. Sediment particle size, organic carbon content, and pore saturation control redox conditions in sediments and thus strongly influence Fe, S, and U biogeochemistry. These findings help to illuminate the mechanistic linkages between hydrology, sediment texture, and biogeochemistry. They further provide enhanced contextual and conceptual underpinnings to support reactive transport modeling of uranium, other contaminants, and nutrients in redox-variable floodplains—a subject of importance to BER research missions. Cyclic redox variability has major implications for mobility of carbon (C), nitrogen (N), and metal contaminants in groundwater and surface waters. Redox-variable, organic-enriched sediments mediate the mobility of C, N, Fe, S, U, and metal contaminants regionally in the UCRB. Organic-enriched sediments were established to regionally mediate groundwater quality within the UCRB. | |||
| 04/07/2017 | Direct Measurement of Protein Dynamics In Vivo | Center for Structural Molecular Biology | Small-Angle Neutron Scattering | Spallation Neutron Source/High Flux Isotope Reactor | Molecular Structure | Quasi-elastic neutron scattering was used to study the protein GroEL in vivo. Protiated GroEL was over-expressed in deuterated Escherichia coli cells by addition of protiated amino acids during induction of GroEL. The data showed retardation factors of ~2 and ~4 for the internal dynamics and global diffusion of the protein, compared to those of the protein in solution at the same concentration. Comparison with literature values suggests that the effective diffusivity of proteins depends on the length or time scale probed. Selective isotope labeling of biomolecules in cells opens up new lines of biological research using “in-cell neutron scattering” to extract information on dynamics of biomolecular systems that is unobtainable by other analysis techniques. Instruments and FacilitiesQuasi-elastic neutron scattering, BASIS neutron backscattering spectrometer at Spallation Neutron Source at Oak Ridge National Laboratory. | |||
| 04/18/2017 | Could This Enzyme Help Turn Biofuel Waste into Something Useful? | Structurally Integrated Biology for the Life Sciences | X-ray Macromolecular Crystallography | Advanced Light Source | Molecular Structure | Joint BioEnergy Institute study targets LigM for its role in breaking down aromatic pollutants A protein used by common soil bacteria is providing new clues in the effort to convert aryl compounds, a common waste product from industrial and agricultural practices, into something of value. The protein structure of the enzyme LigM was determined using X-ray crystallography, revealing novel structural elements (red in figure) and a conserved tetrahydrofolate-binding domain (gray), with LigM binding to its substrates (green) using internal binding cavities. Researchers characterized aryl O-demethylation by LigM in Sphingomonas paucimobilis, a soil bacterium that metabolizes aryl compounds derived from lignin—the stiff, organic material that gives plants their structure. In biofuel production, aryl compounds are a byproduct of the breakdown of lignin, some pathways of which involve demethylation, an often critical precursor to additional modification steps of lignin-derived aryl compounds. The simple, single-enzyme system of LigM, as well as its functionality over a broad temperature range, makes it an attractive demethylase for use in aromatic conversion. Other findings included: half the LigM enzyme was homologous to known structures with a tetrahydrofolate-binding domain that is found in both simple and complex organisms; the other half of LigM’s structure is completely unique, providing a starting point for determining where its aryl substrate-binding site is located; and LigM is a tyrosine-dependent demethylase. This research provides groundwork needed to aid in developing an enzyme-based system for converting aromatic waste into useful products. Instruments and FacilitiesBeamline 8.2.2 and X-ray macromolecular crystallography at the Berkeley Center for Structural Biology, Advanced Light Source, Lawrence Berkeley National Laboratory. | |||
| 07/11/2017 | Brown Rot Fungi Reveal a New Approach for Biomass Conversion to Fuels and Chemicals | Center for Structural Molecular Biology | Small-Angle Neutron Scattering | Spallation Neutron Source/High Flux Isotope Reactor | Molecular Structure | A multimodal approach used in this study examined wood decay by the brown rot fungi, Gloeophyllum trabeum or Rhodonia placenta, that degrade wood using a chelator-mediated Fenton (CMF) reaction. Small-angle neutron scattering (SANS) showed changes in microfibril bundling and lignin structure during biomass breakdown. Complementary approaches, sum frequency generation (SFG) spectroscopy, X-ray diffraction, atomic force microscopy (AFM), and transmission electron microscopy (TEM) also contributed information on nanoscale structural changes in wood over time. Woods studied were southern yellow pine (Pinus spp.) and birch (Betula verrucosa Ehr.). The data support a degradation mechanism in which sugars released by non-enzymatic action diffuse from the cell wall, facilitated by increasing the porosity of the cell walls. This is a paradigm shift in understanding the mechanism of brown rot fungal degradation. Instruments and FacilitiesSmall angle neutron scattering (SANS) – Bio-SANS at High Flux Isotope Reactor (HFIR) at Oak Ridge National Laboratory (ORNL). Sum frequency generation (SFG) spectroscopy, broadband SFG system. Chelator-mediated Fenton treatments (CMF) and cellulase treatment. X-ray diffraction analysis (XRD) – PANalytical Empyrean diffractometer, The Netherlands, equipped with a Cu X-ray source. Attenuated total reflectance Fourier transform infrared analysis (ATR-FTIR): Nicolet 8700 FTIR Spectrometer (Thermo Scientific) equipped with a smart iTR diamond ATR unit, a KBr beam splitter, and a deuterated triglycine sulfate (DTGS) detector. Atomic force microscopy (AFM) of brown-rotted wood surfaces. Nanoscope IIIa AFM-Digital Instruments, Santa Barbara, California with three 5 µm scans. Transmission electron microscopy (TEM) – Philips CM12 TEM instrument (Philips, Eindhoven, The Netherlands); images recorded on Kodak 4489 negative film and the films subsequently scanned using an Epson Perfection Pro 750 film scanner. | |||
| 09/19/2019 | Dynamics on Cellulose Show Two Important Populations from Neutron Scattering and Simulations | Center for Structural Molecular Biology | Small-Angle Neutron Scattering | Spallation Neutron Source/High Flux Isotope Reactor | Molecular Structure | Biomass pretreatment is necessary to make cellulose accessible to hydrolysis for conversion to biofuels. Understanding water’s role in cellulose reactivity will aid discovery of the underlying processes that change biomass morphology and reactivity during different pretreatment regimes for biofuels production. In this study, cellulose-water interactions were examined using neutron scattering supported by molecular dynamics simulation. The data show two distinct populations of water molecules—one tightly “bound” to the surface and the other interfibrillar and translationally mobile. Accurate models of hydration water in the cell wall can address fundamental questions about cellulose-water interactions. The mobility of the interfibrillar water is also important to enzyme and chemical attack and is distinct from the bound and bulk water. Instruments and FacilitiesDeuterium labeling, neutron scattering, and molecular dynamics simulation; quasi-elastic neutron scattering (QENS) using BASIS, the Backscattering Spectrometer, at Oak Ridge National Laboratory (ORNL) Spallation Neutron Source (SNS). Structural characterization using small-angle neutron scattering (SANS) CG-3 Bio-SANS instrument at High Flux Isotope Reactor (HFIR) facility of ORNL and X-ray diffraction (XRD) analysis combined with SANS. Wide-Angle X-ray Diffraction (WAXD) using theta-theta goniometer Bruker D5005 instrument; quasi-elastic neutron scattering (QENS) using BASIS, the Backscattering Spectrometer at ORNL SNS; Molecular Dynamics (MD) Simulations (GROMACS software and the CHARMM C36 carbohydrate force field); and TIP4P water model. | |||
| 11/15/2017 | Cyanobacterial Studies Examine Cellular Structure During Nitrogen Starvation | Center for Structural Molecular Biology | Small-Angle Neutron Scattering | Spallation Neutron Source/High Flux Isotope Reactor | Molecular Structure | Using nondestructive neutron scattering techniques, scientists are examining how single-celled organisms called cyanobacteria produce oxygen and obtain energy through photosynthesis. Collaborators are conducting a series of experiments to study the behavior of phycobilisomes—large antenna protein complexes in cyanobacteria cells. Phycobilisomes harvest light to initiate photosynthesis, and a better understanding of this process could help researchers design more efficient solar panels and other artificial structures that mimic natural systems. Neutrons can analyze these delicate structures without damaging or killing the cyanobacteria and with more spatial accuracy than other techniques like microscopy. Biological small-angle neutron scattering (bio-SANS) allows observing what’s happening at the nanoscale level in real time in a living cell. Phycobilisomes attach to cellular membranes where the light-dependent reactions of photosynthesis take place. Changing the antenna complexes of the phycobilisomes can have dramatic and far-reaching consequences in cyanobacteria. Artificially modifying phycobilisomes by deleting certain genes in the cells caused structural defects in the cellular membranes and other cell physiology, allowing scientists to observe the resulting structural changes. Starving the cyanobacteria for nitrogen naturally modifies the antenna complexes, causing the antenna to decrease in size and leading to significant cellular membrane modifications, because the cells break down the phycobilisomes and use them as an alternative nitrogen source to survive. By determining the extent of these changes, the team hopes to better understand the structure-function relationship between cellular organization and natural modification. These processes can be immediately reversed by restoring nitrogen to the cells. The researchers plan to compare these results to those recorded from their genetic studies to explore the differences between artificial and natural modifications and their effects on the intracellular makeup of cyanobacteria. Instruments and FacilitiesPhotosynthetic Antenna Research Center (PARC), a DOE BES-funded Energy Frontier Research Center based at Washington University at St Louis. Small angle neutron scattering (SANS) was performed at the DOE-BER supported Bio-SANS instrument, beamline CG-3, at Oak Ridge National Laboratory’s High Flux Isotope Reactor. | |||
| 12/13/2017 | Binding of Alternative Substrates to Nitrogenase | Structural Molecular Biology Resource | X-ray Macromolecular Crystallography | Stanford Synchrotron Radiation Lightsource | Molecular Structure | The nitrogenase enzyme, which catalyzes the reduction of atmospheric dinitrogen (N2) to ammonia (NH3) in bacteria and archaea, is an integral component of the global nitrogen cycle. The nitrogen cycle is of interest because nitrogen availability can affect the rate of key ecosystem processes. Any perturbations of the global nitrogen cycle can negatively affect the natural environmental systems and also have a deleterious effect on human health. Fossil fuel combustion, the release of nitrogen in wastewater, and the use of artificial nitrogen fertilizers have dramatically altered the global nitrogen cycle. Therefore, a detailed understanding of the catalytic mechanism of the nitrogenase enzyme, critical to the cycle, is of vital importance. The most abundant nitrogenase, the molybdenum-dependent enzyme, contains two metalloprotein components, molybdenum-iron (MoFe) protein and Fe protein. During the reduction of N2 to NH3, single-electron transfers take place between the donor Fe protein and the MoFe protein. During this process the so-called “P-cluster” [an (8Fe-7S) cluster] in the MoFe protein functions as an intermediate electron carrier between the Fe protein and the active site cofactor (FeMo-co) in the MoFe protein. The nitrogenase enzyme is also capable of reducing small organic substrates such as acetylene (to produce ethylene), and research has demonstrated that N2 binding can inhibit acetylene binding, but acetylene does not necessarily inhibit N2 reduction. Currently, very little is understood about this apparent conundrum since no substrate-bound structures of nitrogenase were known. Certain mutations (in particular at residues Val-70 and Arg-96) in the MoFe protein can result in an increased ability to reduce normally rather poor substrates (including propyne or 1-butyne), while other mutations can decrease the enzyme’s activity against N2 and acetylene. Some MoFe variants can trap alternate substrates (e.g., acetylene or cyanide) in the active site. Researchers at Washington State University have determined the crystal structure of a variant of the MoFe protein with a glutamine residue in place of the native arginine at position 96. The mutant protein was crystallized in the presence of the alternative substrate acetylene, and the structure was solved by molecular substitution to 1.7-Angstrom (Å) resolution using data collected at the Stanford Synchrotron Radiation Lightsource (SSRL) beamline 12-2. The structure shows that the acetylene is trapped in a channel in close proximity to the FeMo-co site (see figure). Complementary theoretical calculations support the observation of acetylene binding at this site and are also consistent with more favorable interactions in the variant MoFe protein compared to the native MoFe protein. This is the first structural evidence of a substrate trapped in the nitrogenase MoFe protein and is consistent with earlier assignments of proposed substrate pathways and substrate binding sites. | |||
| 01/17/2018 | "Zip Code" Mechanism for Hormone Signaling Revealed | Center for BioMolecular Structure | X-ray Macromolecular Crystallography | National Synchrotron Light Source II, Stanford Synchrotron Radiation Lightsource | Molecular Structure | Named after the Greek goddess who spun the thread of life, Klotho proteins play an important role in the regulation of longevity and metabolism. Alpha-Klotho is a membrane-spanning protein expressed predominantly in the kidney, as well as in the brain. Mice lacking a-klotho exhibit a range of signs associated with aging and have elevated blood phosphate levels. Like a-klotho, ß-klotho functions as a co-receptor for endocrine fibroblast growth factors (FGFs). FGF21 is secreted from the liver following fasting, acting in fat cells and the brain to induce metabolic adaptation to fasting and responses to stress. Although FGF receptors (FGFRs) are expressed in a wide range of tissues, expression of the ß-klotho “longevity” protein in the liver, fat, and brain restricts the target organs of these endocrine FGFs. In a recent Yale Medical School-led study, researchers revealed the three-dimensional (3D), high-resolution structure of ß-Klotho, illuminating its intricate mechanism and potential for antiaging therapeutics, as well as for treating a wide range of medical conditions. X-ray crystallography data revealed critical molecular interactions required for hormone binding and cell activation. The specific “zip code”-like interactions of ß-klotho receptors appear to regulate critical metabolic processes in the liver, kidneys, and brain, among other organs. Analysis yielded several insights, including that ß-Klotho is the primary receptor that binds to FGF21, a key hormone that stimulates insulin sensitivity and glucose metabolism, causing weight loss. The researchers believe this new understanding can guide the development of therapies by improving the biological activity of FGF21. Also found were a new variant of FGF21 that has 10 times higher potency and cellular activity and the evidence of how a structurally related enzyme (glycosidase) that breaks down sugars evolved into a receptor for a hormone that lowers blood sugar. Researchers believe the untangled ß-Klotho structure presents a platform for exploring and developing agents that either enhance or block the pathway, enabling therapies for conditions such as liver cancer and bone diseases. | X-ray data were collected from ß-klotho–receptor crystals at Stanford Synchrotron Radiation Lightsource (SSRL), SLAC National Accelerator Laboratory, beamline 14-1 as part of the program to aid National Synchrotron Light Source (NSLS) users prior to NSLS-II operations. | ||
| 01/18/2018 | Structure of a Flavoenzyme Assembly Intermediate | Structural Molecular Biology Resource | X-ray Macromolecular Crystallography | Stanford Synchrotron Radiation Lightsource | Molecular Structure | Enzymes frequently depend on an electron transport cofactor for executing catalytic functions such as reduction-oxidation (redox) reactions. For flavoenzymes, the cofactor is flavin adenine dinucleotide (FAD), whose binding type with the enzyme impacts the redox potential and thus reaction chemistry, such as for metabolism and detoxification. Researchers discovered that the structure of an assembled flavoenzyme intermediate reveals the mechanism of covalent flavin binding in respiration. Assembly factors include SdhAF2 in humans, SdhE in Escherichia coli, and Sdh5 in yeast. Other revelations include that mitochondrial flavoenzymes drive both noncovalent and covalent redox reactions and that the assembly factor (SdhE, a small protein of ~90 to 140 amino acids, conserved in all kingdoms) in the structure of the SdhE:FrdA complex with covalent FAD stabilizes a conformation of the flavoprotein subunit FrdA that favors succinate oxidation. Researchers fixed the E. coli FrdA-SdhE intermediate via site-specific crosslinking, resolving the structure to 2.6 angstroms (Å). This study identified that SdhE stabilizes an FrdA conformation that likely enables the mechanism of autocatalytic covalent flavinylation. FrdA’s FAD-binding domain and capping domain both interact with SdhE, but structural data revealed a 10.8° difference in their angles. The investigators believe that domain rotation affects flavinylation, showing that enzymes are tuned to catalyze reactions in different ways and that conformational diversity can directly relate to catalytic mechanism diversity. | |||
| 12/14/2017 | Using Plants to Immobilize and Stabilize Arsenic in the Soil | Center for BioMolecular Structure, Structural Molecular Biology Resource | X-ray Absorption and Emission Spectroscopy, X-ray Fluorescence Imaging | National Synchrotron Light Source II, Stanford Synchrotron Radiation Lightsource | Chemical and Elemental Information | Phytostabilization is a cost-effective long-term bioremediation technique for the immobilization of metalliferous mine tailings. However, the biogeochemical processes affecting metal(loid) molecular stabilization and mobility in the root zone remain poorly resolved. In this study, the roots of Prosopis juliflora were grown for up to 36 months in compost-amended pyritic mine tailings from a federal Superfund site and then investigated by microscale and bulk synchrotron X-ray absorption spectroscopy and multiple energy micro-X-ray fluorescence imaging to determine iron, arsenic, and sulfur speciation, abundance, and spatial distribution. Two distinct mechanisms of arsenic detoxification were identified: (1) As(V) bound to ferric sulfate plaques on root surfaces and (2) As(III) complexes in root vacuoles. | |||
| 08/30/2018 | Directed Evolution Mimics Allosteric Activation of Enzymes | Structural Molecular Biology Resource | X-ray Macromolecular Crystallography | Stanford Synchrotron Radiation Lightsource | Molecular Structure | Biocatalysis is emerging as a powerful methodology for green, sustainable chemical synthesis. While active-site mutagenesis of nonessential residues is a straightforward way to alter substrate specificity and improve function, residues throughout a protein’s structure can also influence catalytic activity. These remote mutations are reminiscent of allosteric effects, where the binding of a ligand or partner protein (an effector) distant from the active site alters enzyme function. Many, if not most, enzymes are allosterically regulated and are an important source of catalytic diversity. Unfortunately, the removal of the native effector often results in loss of activity and can be a major barrier to the successful implementation of enzymes as biocatalysts. This scenario is exemplified by the enzyme tryptophan synthase, which is a model system for understanding allosteric function as well as a highly desirable biocatalyst for the synthesis of noncanonical amino acids. Nobel Laureate Frances Arnold (California Institute of Technology) and colleagues solved the high-resolution crystal structures of the (1) wild-type enzyme, (2) an intermediate in the lineage, and (3) the final variant, all using the Stanford Synchrotron Radiation Lightsource (SSRL) beamline 12-2. The structures revealed that the activating mutations have only minor structural effects on their immediate environment; however, they stabilize the large-scale motion of a subdomain to favor an otherwise transiently populated, closed conformational state (see figure). This increase in stability enabled the first structural description of tryptophan covalently bound in catalytically active tryptophan synthase, confirming key features of catalysis. These data combine to show that sophisticated models are not a prerequisite to mimicking the activation via directed evolution, opening the way to engineering stand-alone versions of diverse allosteric enzymes. | |||
| 08/21/2018 | Potent Bacteria | Structural Biology Center | X-ray Macromolecular Crystallography | Advanced Photon Source | Molecular Structure | A collaboration of scientists from the U.S. Department of Energy’s (DOE) Argonne National Laboratory has found a special strain of soil bacteria that protects itself from other organisms by producing highly toxic compounds and confining them to a “molecular dungeon.” Unusual properties of the enediyne compound include acting as powerful antitumor agent or antibiotic, but the researchers discovered that this family of soil bacteria also produces a tightly bound protein that sequesters the toxic compound from the rest of the organism. The study has implications for cancer treatment and understanding how human cancer cells develop resistance to natural product–based chemotherapies. | |||
| 11/15/2018 | Controlled Self-Assembly of a Designed Nucleoprotein | Structural Molecular Biology Resource | Solution X-ray Scattering | Stanford Synchrotron Radiation Lightsource | Molecular Structure | Nature employs a limited set of basic building blocks such as nucleic acids and proteins to create strikingly diverse materials and machines that perform their function with very high fidelity. Inspired by this, synthetic biology and bio-nanotechnology research aims to develop self-assembled structures and materials from the same biological building blocks with functions compatible with or complementary to natural ones that ultimately can surpass those produced by evolution. In this study, the authors developed a synthetic nucleoprotein assembly that combines a four-helix bundle (RIDC3) with a 10 base pair single-stranded DNA sequence (RIDC3-10a) and its complementary sequence (RIDC3-10b). The construct combines three prominent classes of intermolecular interactions (Watson-Crick base pairing, DNA-protein interactions, and protein-metal coordination with Zn2+) to self-assemble with high structural order and specificity in a manner reminiscent of natural nucleoproteins like the ribosome. The different structures obtained under a variety of conditions were characterized and structurally identified using equilibrium and in situ time-resolved small-angle X-ray scattering (SAXS) measurements at the Stanford Synchrotron Radiation Lightsource (SSRL) beamline 4-2, electron microscopy, and molecular dynamics. The analysis showed that the RIDC3-DNA crystals were discrete two-dimensional (2D) molecular layers that stacked up in 3D (see figure). While the modular nature of such multicomponent systems should offer distinct advantages in the construction of structurally tunable materials, the intricate architecture of the RIDC3-DNA assembly also highlights the opportunities and challenges inherent in designing artificial nucleoprotein complexes that arise from the distinct structural and chemical properties of proteins and nucleic acids. | |||
| 02/05/2019 | Peptide Induced Lateral Segregation of Lipids in a Biomembrane | Center for Structural Molecular Biology | Small-Angle Neutron Scattering | Spallation Neutron Source/High Flux Isotope Reactor | Molecular Structure | This study shows, for the first time, that AMPs can disrupt membrane function by inducing lateral segregation of lipids in the bilayer. Previously, AMPs were generally thought to disrupt membranes by forming transmembrane pores. This study provides fundamental insights into AMP-membrane interactions and new mechanistic insights into their mode-of-action. | Small-angle neutron scattering (SANS) with contrast matching showed that the short anti-microbial peptide (AMP), Aurein 1.2, induced lateral segregation of lipids in a biomembrane at concentrations that are toxic to bacteria. Aurein induces significant lateral segregation in an initially uniform lipid bilayer composed of zwitterionic lipid and anionic lipid. QENS showed lipid lateral motion in the fluid phase was reducedeven at low aurein concentrations, providing a cause for the heterogeneity. Work was performed at the DOE BER-supported Bio-SANS instrument at HFIR and the BES-supported BASIS instrument at SNS. |
| |
| 02/25/2019 | Breakthrough Shines Light on Disease-Fighting Protein | Stanford-SLAC Cryo-EM Center, Structural Biology Center | Cryo-Electron Microscopy, X-ray Macromolecular Crystallography | Advanced Photon Source, Stanford Synchrotron Radiation Lightsource | Molecular Structure | A team of researchers have obtained the highest-resolution structure of the fungal protein Hsp104, which could be used to hinder the formation of certain degenerative diseases. Using the Advanced Photon Source at Argonne National Laboratory, the researchers combined X-ray crystallography and cryo-electron microscopy (cryo-EM) to also verify that these protein-formed hexamers, once believed flat, have a helical structure. Known as a chaperone, the hexameric AAA+ protein Hsp104 helps in the natural folding processes of proteins for proper cell function and may be able to repair misfolded or aggregated proteins that can lead to protein-caused abnormalities. | |||
| 02/18/2019 | An Efficient and Economical Strategy for the Design of Functional Metalloproteins | Structural Molecular Biology Resource | X-ray Macromolecular Crystallography | Stanford Synchrotron Radiation Lightsource | Molecular Structure | Although numerous strategies have been used to create new metalloproteins, pre-existing knowledge of the tertiary and quaternary protein structure is often required to generate suitable platforms for robust metal coordination and activity. F. A. Tezcan (University of California, San Diego) and colleagues recently reported an alternative and easily-implemented approach using metal active sites by covalent tethering, in which folded protein building blocks are linked by a single disulfide bond to create diverse metal coordination environments within novel protein-protein interfaces (MASCoT). Metalloproteins generated using this method uniformly bind a wide array of first-row transition metal ions [manganese (Mn+2), iron (Fe+2), cobalt (Co+2), nickel (Ni+2), copper (Cu+2), and zinc (Zn+2)] with physiologically relevant thermodynamic affinities. The researchers were able to create two new metalloproteins with stable Mn binding sites. Structures were obtained using X-ray crystallography data collected on Stanford Synchrotron Radiation Lightsource (SSRL) beamline 12-2 using Mn multi-wavelength anomalous diffraction (MAD) phasing (see figure). The Mn coordination sphere in Mn-CH2E includes (1) three meridional histidine side chains and (2) a single glutamate that completes a square-planar ligand arrangement around a trans-(OH2)2Mn+2 unit. These aquo ligands are in turn engaged in strong hydrogen-bonding interactions with the two other engineered histidine and glutamate side chains. By contrast, the Mn+2 coordination sphere determined for Mn–CH2EY includes all four designed histidine residues and a bound glutamate, collectively reminiscent of the nonheme Fe site found in the photosynthetic reaction center of Rhodobacter sphaeroides. New metalloproteins were also created with other metals that could, for example, bind small gaseous molecules. The fact that all of these functional features would be difficult to design on their own but are simultaneously achieved through MASCoT with minimal engineering attests to the expediency of the covalent tethering strategy. Since this strategy is predicated on the use of well-folded protein building blocks and natural amino acids, its application in the laboratory evolution of enzymatically active metalloproteins involved in oxidative and hydrolytic processes can be realized. It may become an integral part of synthetic biology approaches. | |||
| 02/22/2019 | DNA Gets a Bigger Alphabet | Structural Molecular Biology Resource | X-ray Macromolecular Crystallography | Stanford Synchrotron Radiation Lightsource | Molecular Structure | Synthetic biology has made available a mutable genetic system, built from eight different building blocks, with potential use spanning creating novel proteins to pursuing extra-terrestrial life. | Four new DNA bases were synthesized and combined with the four natural bases to give an expanded genetic Hachimoji alphabet, capable of folding into a standard double helix and producing viable RNA. |
| |
| 03/05/2019 | Understanding Cadmium Distribution in Hyperaccumulating Plants for Efficient Phytoremediation | Structural Molecular Biology Resource | X-ray Fluorescence Imaging | Stanford Synchrotron Radiation Lightsource | Chemical and Elemental Information | Cadmium (Cd) is a toxic metal that contaminates soils, particularly in industrialized regions. It can threaten human health, particularly because it readily accumulates in the edible parts of crops, thereby exposing humans through their diet. Some particular plant species are known to be hyperaccumulators, taking up and storing much more Cd (up to 100 mg/kg) than is typical. This characteristic can be exploited to remove Cd from soil, a process termed phytoremediation. Metal hyperaccumulation has traditionally been attributed to rapid uptake by roots, enhanced transport of metals from roots to shoots, and, finally, efficient metal sequestration and detoxification in the shoots. However, it is not well understood how and whether metals can be redistributed throughout the plant as it ages. The current work sought to investigate whether Cd could be transported in phloem, a tissue in vascular plants that is responsible for moving nutrients around to parts of the plant where they are needed. Nutrients are often redistributed from old to developing tissues during plant growth, but, before this study, it was unclear whether the same process occurs for toxic metals. Hu et al. used X-ray fluorescence (XRF) imaging performed on beamline14-3 at the Stanford Synchrotron Radiation Lightsource (SSRL), in combination with bulk measurements of the Cd concentration in both old and young leaves and stems, to determine whether Cd would be transported from old to young parts of the plant in the hyperaccumulator Sedum alfredii. Their results show that Cd concentrations were in fact significantly higher in young, developing leaves and stems relative to mature ones. Cross sections of the stems also showed that the distribution of Cd among different types of tissues varied between the young and old stems. In younger stems, Cd was found in the pith and cortex. In the older stems, tissues near the junction of leaves and stems showed elevated Cd presence. Additionally, Cd was elevated in the curvilinear outer layers of the vascular bundles, where phloem tissue is located. In summary, Hu et al. demonstrated the ability of the hyperaccumulating plant S. alfredii to reallocate Cd from old to young tissues via translocation in phloem. Reallocation of Cd among different tissues may contribute to the survival of the plant. Although hyperaccumulators have high tolerances toward metals, the redistribution of Cd may ensure that Cd concentration does not become too high at any location as the plant ages. Finally, redistribution of Cd has the benefit that less Cd is lost in senesced leaves, enhancing the efficiency of phytoremediation. | |||
| 04/05/2019 | Influence of Chemically Disrupted Photosynthesis on Cyanobacterial Thylakoid Dynamics | Center for Structural Molecular Biology | Small-Angle Neutron Scattering | Spallation Neutron Source/High Flux Isotope Reactor | Molecular Structure | This study supports the hypothesis that the proton pressure buildup within the lumenal space during photosynthesis drives dynamical changes in the cyanobacterial thylakoids, that are directly and quantitatively correlated with the mechanical properties of photosynthetic membranes. | Herbicide treatment effects on the dynamical behavior of cyanobacterial photosynthetic membranes (thylakoids) were measured. Changes in membrane mechanical properties were correlated with their structural organization. This provides an important step in understanding the flexibility of the thylakoids and their active function during energy conversion. |
| |
| 04/12/2019 | What Enables the Fastest Carbon Dioxide Fixation by Reductive Carboxylase? | Structural Molecular Biology Resource | X-ray Macromolecular Crystallography | Stanford Synchrotron Radiation Lightsource | Molecular Structure | Persistent challenges in atmospheric CO2 capture and conversion have stimulated an increasing interest in understanding and exploiting CO2-fixation mechanisms. Enoyl-CoA carboxylases/reductases (ECRs) are the most efficient CO2-fixing enzymes found in nature to date, outcompeting RuBisCO (the key enzyme in photosynthesis) by more than an order of magnitude in activity. However, the molecular mechanisms underlying ECR’s extraordinary catalytic activity remain elusive. In a large consortium that included Tobias Erb (Max Planck Institute for Terrestrial Microbiology) and scientists at the U.S. Department of Energy’s (DOE) SLAC National Accelerator Laboratory (SLAC) and the DOE Joint Genome Institute, multiple crystallographic approaches were used, including synchrotron and X-ray free electron laser (XFEL) experiments, to study the fixation mechanism of ECR from the actinobacterium Kitasatospora setae. The researchers used the Stanford Synchrotron Radiation Lightsource (SSRL) beamline 12-2 to collect data from a ternary structure containing ECR and bound butyryl-CoA and NADPH. The complex formed a tetramer organized as a dimer of dimers with both “open” and “closed” states with respect to the active site. Dimer subunits are stabilized with the binding of NADPH. Two of these subunits, one open and one closed, form the overall dimer. This tetramer that differentiates into a dimer of dimers of open- and closed-form subunits suggests that the enzyme operates with “half-site reactivity” and in conformational synchrony to achieve the observed high catalytic rates. | |||
| 05/03/2019 | New Approach for Solving Protein Structures from Tiny Crystals | Center for BioMolecular Structure | X-ray Macromolecular Crystallography | National Synchrotron Light Source II | Molecular Structure | Using X-rays to reveal the atomic-scale three-dimensional structures of proteins has led to countless advances in understanding how these molecules function in bacteria, viruses, plants, and humans. To grow some proteins into crystals large enough for this approach, scientists at the U.S. Department of Energy’s (DOE) Brookhaven National Laboratory (BNL) and colleagues at Columbia University developed an approach for solving protein structures from tiny crystals. To examine previously inaccessible microcrystals, they are using unique sample-handling, signal-extraction, and data-assembly approaches, along with a beamline capable of focusing intense X-rays at Brookhaven National Laboratory’s National Synchrotron Light Source II. | |||
| 07/10/2019 | Structuring Sweetness: What Makes Stevia 200 Times Sweeter than Sugar? | Structural Biology Center | X-ray Macromolecular Crystallography | Advanced Photon Source | Molecular Structure | The major ingredient in the product Stevia, rebaudioside A (called “RebA”), has been published, enabling a determination of the three-dimensional structure of its proteins. Along with the mostly known genes and proteins in the biochemical pathway, there is potential for manufacturing noncaloric products without the aftertaste some associate with the intensely sweet Stevia. Researchers used X-ray crystallography capabilities at the Advanced Photon Source’s Structural Biology Center at Argonne National Laboratory to determine RebA’s structure and how the key plant enzyme builds high-intensity sweetness. | |||
| 07/10/2019 | Structural Characterization of Chitinases from <i>Agave tequilana</i> | Structural Molecular Biology Resource | Solution X-ray Scattering | Stanford Synchrotron Radiation Lightsource | Molecular Structure | The economically important plant genus Agave is cultivated to produce fiber, food, beverages, and fuel. Within this genus, A. tequilana is primarily used for the elaboration of beverages and in soil conservation, but research indicates it’s potential as a source of lignocellulosic bioenergy feedstocks. It therefore has great socioeconomic and agroecological value, especially in hot and drought-prone regions of the world. Despite its adaptability, these plants are subject to diseases such as those caused by insects and fungus. Plant chitinases are enzymes that hydrolyze chitin, a molecule present in fungal cell walls, most insect exoskeletons, yeasts and algae, crustacean shells, and other organisms. They are also involved in numerous plant physiological events such as abiotic stress responses, including antifungal activity and defense mechanisms. In this study, the authors describe the solution structure and biophysical characterization of two chitinases from A. tequilana as determined by small-angle X-ray scattering (SAXS) at the Stanford Synchrotron Radiation Lightsource in combination with theoretical structure prediction using Robetta, a protein structure prediction service. The low-resolution structures both exhibited two distinct domains connected by a linker region, and the theoretically predicted Rosetta structure showed consistency with the solution structures and could be docked into the SAXS envelopes. Interestingly, AtChi1 and AtChi2 both showed antifungal activities, suggesting that the Agave enzymes are involved in host defense mechanisms. Therefore, these enzymes could be considered as pathogenesis-related proteins with an important role in the plant’s defense mechanism against fungi. | |||
| 11/19/2019 | How Essential Membrane Lipids Interact to Regulate Cellular Processes | Structural Biology Center | X-ray Macromolecular Crystallography | Advanced Photon Source | Molecular Structure | The regulation of many cellular processes relies on interactions between sphingomyelin and cholesterol, two essential lipids in the cell’s plasma membrane. Researchers collected diffraction data at the U.S. Department of Energy’s Advanced Photon Source (APS) and examined the three-dimensional molecular interaction between the lipid-binding protein Ostreolysin A and sphingomyelin/cholesterol complexes. The results improved current understanding of how these lipids interact to carry out regulatory functions vital for controlling many signaling processes within the cell such as regulating cholesterol synthesis and uptake. This is important because the stability and integrity of the plasma membrane depend on proper levels of cholesterol; the study will instruct further research, but the structural-level mechanisms of interaction remain poorly understood. | |||
| 11/29/2019 | Ammonia-Salt Solvent Promotes Cellulosic Biomass Deconstruction Under Ambient Processing Conditions | Center for Structural Molecular Biology | Small-Angle Neutron Scattering | Spallation Neutron Source/High Flux Isotope Reactor | Molecular Structure | A:At-pretreated cellulose requires ∼50-fold less enzyme for saccharification into fermentable sugars for biorefining. The reaction takes place under ambient conditions, requiring much less energy input compared to most conventional thermochemical pretreatment processes. | A new highly efficient ammonia:ammonium salt (A:At) solvent-based biomass pretreatment was devised. Using time-resolved small-angle neutron scattering, it was possible to determine the mechanism of dissolution of crystalline cellulose into a “molecular” solution. Complementary molecular dynamics simulations showed how the A:At solvent disrupted the cellulose hydrogen bonding network. | Time-resolved small-angle neutron scattering (SANS) studies performed in deuterated A:At solvents revealed the structural changes in cellulose as they occurred. Enzyme digestion studies quantified the efficiency of the reactions. A:At solvent allowed 80-85% recovery of lignin from corn stover under ambient pretreatment conditions. | |
| 12/09/2019 | Research at the Advanced Photon Source Leads to New Ebola Drug | Structural Biology Center | X-ray Macromolecular Crystallography | Advanced Photon Source | Molecular Structure | The ability to examine how specific antibodies react can lead to treatments for deadly diseases and often depends on determining how proteins in our bodies behave. Research at Argonne National Laboratory’s Advanced Photon Source enables these insights and has led to the development of a promising drug for Ebola. Using specialized beamlines, researchers revealed how two human antibodies were able to neutralize the virus by examining single-crystal samples of a complex combining the Ebola protein with the protective antibodies. A drug based on one of these antibodies proved remarkably effective during a recent Ebola outbreak and clinical trials in the Democratic Republic of Congo. | |||
| 08/29/2020 | How Proteins Remodel DNA in Bacteria Under Stress | National Center for X-Ray Tomography, Structurally Integrated Biology for the Life Sciences | Soft X-ray Tomography, Solution X-ray Scattering, X-ray Macromolecular Crystallography | Advanced Light Source | Molecular Structure, Cell and Tissue Structure | When bacteria are put in different environments, they can begin to quickly adapt because the proteins that make up their chromosomes can pack and unpack rapidly, regulating gene expression. In bacteria, the proteins responsible for DNA packing are called HU proteins. To examine the DNA packing process under various conditions, researchers visualized the interactions between DNA and HU from Escherichia coli at the micro-, meso-, and nanoscales using soft x-ray tomography, small-angle x-ray scattering, and protein crystallography. | |||
| 08/13/2020 | Cell Membrane Proteins Imaged in 3D | Center for BioMolecular Structure, Structural Biology Center | X-ray Fluorescence Imaging, X-ray Macromolecular Crystallography | Advanced Photon Source, National Synchrotron Light Source II | Chemical and Elemental Information | Using ultrabright X-rays, researchers at the National Synchrotron Light Source II have demonstrated a new technique for imaging proteins in three dimensions with nanoscale resolution. Their approach, which employs X-ray fluorescence microscopy, enables scientists to identify the precise location of proteins within individual cells, reaching the resolution of the cell membrane and the smallest subcellular organelles. Watch a 3-D X-ray movie of a single E.coli cell imaged at NSLS-II with a sub-15 nm X-ray beam, which is the highest resolution X-ray fluorescence tomogram of a biological cell ever collected (as of the paper’s publication date in 2020). Facilities and InstrumentsTwo-dimensional X-ray fluorescence microscopy was performed at the Bionanoprobe (BNP) at beamline 21-ID-D (now at 9-ID-B) at the Advanced Photon Source at Argonne National Laboratory. Three-dimensional X-ray nanotomography was performed at the Hard X-ray Nanoprobe beamline 3-ID at the National Synchrotron Light Source II (NSLS-II) at Brookhaven National Laboratory. | |||
| 07/21/2020 | Role of Solvents in Efficient Biomass Deconstruction | Center for Structural Molecular Biology | Small-Angle Neutron Scattering | Spallation Neutron Source/High Flux Isotope Reactor | Molecular Structure | Direct experimental and computational evidence of a simple physical chemical principle that explains the success of mixing an organic co-solvent, tetrahydrofuran, with water to overcome this recalcitrance. The hydrophilic and hydrophobic biomass surfaces are solvated by single-component nanoclusters of complementary polarity. | ObjectiveUnderstand the effect of tetrahydrofuran (THF):water pretreatment on the nanoscale architecture of biomass and the role these co-solvents play in solubilizing lignin and cellulose. ApproachIn-situ small-angle neutron scattering (SANS) with contrast variation and molecular dynamics (MD) simulations were performed to characterize the biomass structure and the interactions of solvents with biomass components. |
| |
| 02/14/2020 | How Dinosaur Blood Vessels Preserve Through the Ages | Berkeley Synchrotron Infrared Structural Biology Imaging Program | Synchrotron Infrared Hyperspectral Imaging | Advanced Light Source | Chemical and Elemental Information | A team of scientists at the University of Wisconsin–Stout used infrared and X-ray imaging and spectromicroscopy to demonstrate how soft tissue structures may be preserved in dinosaur bones. Performed at Lawrence Berkeley National Laboratory’s (LBNL) Advanced Light Source (ALS), the research countered scientific dogma that protein-based body parts cannot survive more than 1 million years. Samples from a 66-million-year-old Tyrannosaurus rex tibia provided evidence that vertebrate collagen and elastin structures (from blood vessels) may persist across geologic time through two natural processes called Fenton chemistry and glycation. | |||
| 06/23/2020 | Novel Cell Membrane Model Could Be Key to Uncovering New Protein Properties | Center for BioMolecular Structure, Center for Structural Molecular Biology | Cryo-Electron Microscopy, Small-Angle Neutron Scattering, Solution X-ray Scattering | National Synchrotron Light Source II, Spallation Neutron Source/High Flux Isotope Reactor | Molecular Structure, Cell and Tissue Structure | Raft-like bicelles developed in this study provide a novel model membrane system for membrane proteins that require sphingomyelin and cholesterol for proper structure and function. | Bicelles rich in sphingomyelin and cholesterol have been developed to mimic raft-like membranes for membrane protein research. Human amyloid precursor protein C99 in the raft-like bicelles is found to be low-order oligomer in contrast to previous studies. |
| |
| 10/07/2020 | "Missing Link" in Evolutionary History of the Carbon-Fixing Protein Rubisco | Structurally Integrated Biology for the Life Sciences | Solution X-ray Scattering | Advanced Light Source | Molecular Structure | Researchers have discovered a unique, ancient version of Rubisco, the most abundant enzyme on Earth and critical to life as we know it. Found in previously unknown environmental microbes, the newly identified Rubisco lineage, called form I’ Rubisco, provides insight into the evolution of the photosynthetic organisms that underlie global food chains. Contributing to this discovery were scientists at the Structurally Integrated Biology for the Life Sciences (SIBYLS) beamline 12.3.1, located at Lawrence Berkeley National Laboratory’s Advanced Light Source. By analyzing size-exclusion chromatography/small-angle X-ray scattering (SEC-SAXS) data collected at SIBYLS, the team was able to capture how the enzyme’s structure changes during different states of activity. |

