Detergent-Free Method for Photosystem I Stabilization in Structural Studies
10/01/2022
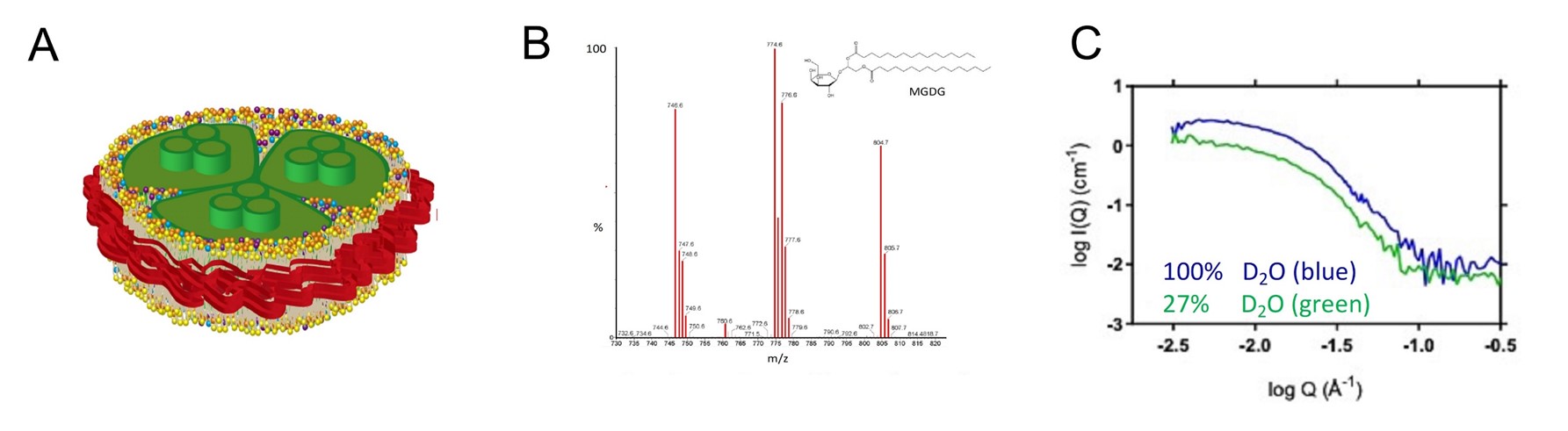
(A) A schematic illustration of photosystem I (green) from a cyanobacterium embedded in a nanodisc (SMALP) to preserve native form and function during in vitro analysis. (B) A mass-spectrometry profile of lipids contained in a cyanobacterial thylakoid membrane, including monogalactosyldiacylglycerol (MGDG). (C) Small-angle neutron scattering analysis of PSI-SMALP at different concentrations of heavy water (D2O) reveal a styrene maleic acid (SMA) copolymer belt (red) around PSI-SMALP. [Reprinted from BBA Bioenergetics, 1863(7):148596, Brady et al. Small angle neutron scattering and lipidomic analysis of a native, trimeric PSI-SMALP from a thermophilic cyanobacteria. Copyright 2022, with permission from Elsevier.]
The Summary
Oxygenic photosynthesis is the process by which plants, algae, and cyanobacteria convert sunlight into chemical energy, fueling all life on earth. It is performed by four membrane-associated protein complexes working in concert: photosystem II (PSII), cytochrome b6f, photosystem I (PSI), and ATP synthase.
The oligomeric states of the individual domains of these protein complexes are relatively similar across kingdoms, except for PSI. PSI exists as a trimer in cyanobacteria and as a monomer in plants and algae. More recently, a tetrameric form has been found in heterocyst-forming cyanobacteria. This dynamic evolutionary history has spurred questions about how structural differences affect the overall function or activity of PSI in vivo. Further, detergent-based methods used to extract PSI from its thylakoid membrane environment for in vitro studies may alter its activity by modifying PSI interactions with thylakoid lipids that determine its function and structural integrity.
Researchers tested an alternative to detergent solubilization to isolate trimeric PSI from the cyanobacterium Thermosynechococcus elongatus in which styrene maleic acid (SMA) copolymers were used to produce membrane protein-containing nanodics—model membranes for solubilizing and studying membrane proteins. The nanodics, called SMA lipid particles (SMALPs), retained the native lipid environment and preserved native protein function.
Small-angle X-ray scattering (SAXS) and neutron scattering (SANS) were used to determine the size and shape of the particles in their fully solvated state. The particle’s proteolipid core and detergent shell or copolymer belt were interrogated separately using contrast variation, a capability unique to SANS. At ~1.5 MDa, PSI-SMALP is the largest SMALP yet isolated. SANS with contrast variation showed, for the first time, a ~1 nm SMA copolymer belt surrounding PSI, explaining the increased energy transfer observed in PSI embedded in SMALPs compared to detergent micelles.
SMA copolymers hold great promise for stabilization of membrane proteins for structural studies. This study provides a structural basis for the observed increase in energy transfer and charge separation in PSI-SMALPs compared to detergent-isolated PSI complexes, highlighting the importance of a native lipid environment for maintaining PSI activity in vitro.
Funding
This work is supported by the Center for Structural Molecular Biology at Oak Ridge National Laboratory (ORNL) and the Biological and Environmental Research Program in the Department of Energy’s Office of Science. The research used resources at the High Flux Isotope Reactor and Spallation Neutron Source, a U.S. Department of Energy Office of Science User Facility operated by ORNL. ORNL is operated by UT-Battelle, LLC, under Contract No. DE-AC05-00OR22725 with the U.S. Department of Energy.
Related Links
References
Brady et al. 2022. Small angle neutron scattering and lipidomic analysis of a native, trimeric PSI-SMALP from a thermophilic cyanobacteria. BBA Bioenergetics, 1863(7):148596. [DOI: 10.1016/j.bbabio.2022.148596]
